
Rice Science ›› 2019, Vol. 26 ›› Issue (6): 343-355.DOI: 10.1016/j.rsci.2018.12.007
• Review • Previous Articles Next Articles
Received:2018-10-12
Accepted:2018-12-19
Online:2019-11-28
Published:2019-08-19
Saiful Islam Md. Sensing and Uptake of Nitrogen in Rice Plant: A Molecular View[J]. Rice Science, 2019, 26(6): 343-355.
Add to citation manager EndNote|Ris|BibTeX
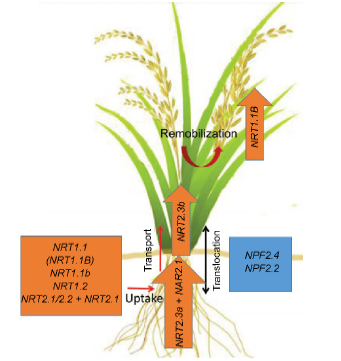
Fig. 1. Candidate NO3- transporters for improving nitrogen use efficiency (NUE) and yield of rice.The orange box indicates genes that could give positive effects on rice for uptake, transport and remobilization of NO3- in normal growth conditions. As examples, NRT1.1 (and another allele, NRT1.1B), NRT1.1b, NRT1.2, NRT2.1 and NRT2.2 (cooperating with NAR2.1) are responsible for NO3- uptake from soil. NRT2.3a (cooperating with NAR2.1) is a root-to-shoot NO3- transporter, and NRT2.3b is responsible for transporting of NO3- to the shoot and remobilizing N into the grain. The blue box indicates genes that can increase rice NUE and yield under salinity, such as NPF2.4 and NPF2.1, by regulating NO3- and K+ distributions in rice plants.
| Gene | Gene locus | Function | Spatial expression | Transcriptional regulation by N | Effect on plant development | Reference |
|---|---|---|---|---|---|---|
| OsAMT1.1 | LOC_Os04g43070 | Dual-affinity NH4+ transporter, root-to-shoot transporter | Root epidermis and vascular tissues, root-shoot junction, vascular bundle, and leaf mesophyll cells | Induced by NH4+ and repressed by N starvation | Promote root growth and grain yield | |
| OsAMT1.2 | LOC_Os02g40710 | NH4+ transporter | Vascular tissues, endodermis and pericycle in primary root tip | Induced by NH4+ and repressed by N starvation | Unknown | |
| OsAMT1.3 | LOC_Os02g40730 | NH4+ transporter | Root vascular tissue and lateral root primordia | Repressed by NH4+ and induced by N starvation | Promote root elongation | |
| OsAMT2.1 | LOC_Os05g39240 | Low-affinity NH4+ transporter | Root, shoot, leaf blade and sheath | Irresponsive to N supply | Unknown | |
| OsAMT2.2 | LOC_Os01g61510 | NH4+ transporter | Unknown | Unknown | Unknown | |
| OsAMT2.3 | LOC_Os01g61550 | NH4+ transporter | Unknown | Unknown | Unknown | |
| OsAMT3.1 | LOC_Os01g65000 | Putative NH4+ transporter | Unknown | Induced by N starvation | Unknown | |
| OsAMT3.2 | LOC_Os03g62200 | Putative NH4+ transporter | Unknown | Unknown | Unknown | |
| OsAMT3.3 | LOC_Os02g34580 | Putative NH4+ transporter | Unknown | Unknown | Unknown | |
| OsAMT4 | LOC_Os03g53780 | Putative NH4+ transporter | Unknown | Unknown | Unknown | |
| OsNRT1.1b | LOC_Os10g40600 | Dual-affinity NO3- transporter | Root epidermis, hairs and vascular tissues, and leaf sheath, leaf blade and culm | Induced by NO3- | Grain yield increase | |
| OsNPF2.4 | LOC_Os03g48180 | Low-affinity NO3- transporter, root-to-shoot transporter | Root epidermis, xylem parenchyma, and phloem companion cells and leaf phloem | Induced by high NO3- supply | Promote root growth and biomass, and grain yield | |
| OsNRT2.1 | LOC_Os02g02170 | High-affinity NO3- transporter | Abundantly expressed in most cell types in root, but weakly expressed in shoot | Induced by NO3- and repressed by NH4+ | Unknown | |
| OsNRT2.2 | LOC_Os02g02190 | High-affinity NO3 transporter | Abundantly expressed in most cell types in roots, but weakly expressed in shoot | Induced by NO3- and repressed by NH4+ | Unknown | |
| OsNRT2.3a | LOC_Os01g50820 | High-affinity NO3- transporter, transport nitrate from root to shoot, not root nitrate uptake | Vascular tissues at root and root-shoot junction | Induced by NO3- and repressed by NH4+ | Promote root growth and biomass | |
| OsNRT2.3b | LOC_Os01g50820 | High-affinity NO3- transporter, sensing cytosolic pH changes | Vascular tissue, mainly in phloem in both root and shoot | Irresponsive to N supply | Improved grain yield and nitrogen use efficiency | |
| OsNRT2.4 | LOC_Os01g36720 | Putative high-affinity NO3- transporter | Lateral root primordia and shoot vascular tissues | Induced by NO3- and repressed by NH4+ | Unknown | |
| OsNAR2.1 | LOC_Os02g38230 | Partner proteins of OsNRT2.1, OsNRT2.2 and OsNRT2.3a | Epidermis and vascular tissue in root and shoot | Induced by NO3- and repressed by NH4+ | Promote root and shoot growth | |
| OsNAR2.2 | LOC_Os04g40410 | Putative partner protein of OsNRT2s | Unknown | Induced by high NO3- and NH4+ | Unknown |
Table 1 Different genes involved in N transport in rice plant.
| Gene | Gene locus | Function | Spatial expression | Transcriptional regulation by N | Effect on plant development | Reference |
|---|---|---|---|---|---|---|
| OsAMT1.1 | LOC_Os04g43070 | Dual-affinity NH4+ transporter, root-to-shoot transporter | Root epidermis and vascular tissues, root-shoot junction, vascular bundle, and leaf mesophyll cells | Induced by NH4+ and repressed by N starvation | Promote root growth and grain yield | |
| OsAMT1.2 | LOC_Os02g40710 | NH4+ transporter | Vascular tissues, endodermis and pericycle in primary root tip | Induced by NH4+ and repressed by N starvation | Unknown | |
| OsAMT1.3 | LOC_Os02g40730 | NH4+ transporter | Root vascular tissue and lateral root primordia | Repressed by NH4+ and induced by N starvation | Promote root elongation | |
| OsAMT2.1 | LOC_Os05g39240 | Low-affinity NH4+ transporter | Root, shoot, leaf blade and sheath | Irresponsive to N supply | Unknown | |
| OsAMT2.2 | LOC_Os01g61510 | NH4+ transporter | Unknown | Unknown | Unknown | |
| OsAMT2.3 | LOC_Os01g61550 | NH4+ transporter | Unknown | Unknown | Unknown | |
| OsAMT3.1 | LOC_Os01g65000 | Putative NH4+ transporter | Unknown | Induced by N starvation | Unknown | |
| OsAMT3.2 | LOC_Os03g62200 | Putative NH4+ transporter | Unknown | Unknown | Unknown | |
| OsAMT3.3 | LOC_Os02g34580 | Putative NH4+ transporter | Unknown | Unknown | Unknown | |
| OsAMT4 | LOC_Os03g53780 | Putative NH4+ transporter | Unknown | Unknown | Unknown | |
| OsNRT1.1b | LOC_Os10g40600 | Dual-affinity NO3- transporter | Root epidermis, hairs and vascular tissues, and leaf sheath, leaf blade and culm | Induced by NO3- | Grain yield increase | |
| OsNPF2.4 | LOC_Os03g48180 | Low-affinity NO3- transporter, root-to-shoot transporter | Root epidermis, xylem parenchyma, and phloem companion cells and leaf phloem | Induced by high NO3- supply | Promote root growth and biomass, and grain yield | |
| OsNRT2.1 | LOC_Os02g02170 | High-affinity NO3- transporter | Abundantly expressed in most cell types in root, but weakly expressed in shoot | Induced by NO3- and repressed by NH4+ | Unknown | |
| OsNRT2.2 | LOC_Os02g02190 | High-affinity NO3 transporter | Abundantly expressed in most cell types in roots, but weakly expressed in shoot | Induced by NO3- and repressed by NH4+ | Unknown | |
| OsNRT2.3a | LOC_Os01g50820 | High-affinity NO3- transporter, transport nitrate from root to shoot, not root nitrate uptake | Vascular tissues at root and root-shoot junction | Induced by NO3- and repressed by NH4+ | Promote root growth and biomass | |
| OsNRT2.3b | LOC_Os01g50820 | High-affinity NO3- transporter, sensing cytosolic pH changes | Vascular tissue, mainly in phloem in both root and shoot | Irresponsive to N supply | Improved grain yield and nitrogen use efficiency | |
| OsNRT2.4 | LOC_Os01g36720 | Putative high-affinity NO3- transporter | Lateral root primordia and shoot vascular tissues | Induced by NO3- and repressed by NH4+ | Unknown | |
| OsNAR2.1 | LOC_Os02g38230 | Partner proteins of OsNRT2.1, OsNRT2.2 and OsNRT2.3a | Epidermis and vascular tissue in root and shoot | Induced by NO3- and repressed by NH4+ | Promote root and shoot growth | |
| OsNAR2.2 | LOC_Os04g40410 | Putative partner protein of OsNRT2s | Unknown | Induced by high NO3- and NH4+ | Unknown |
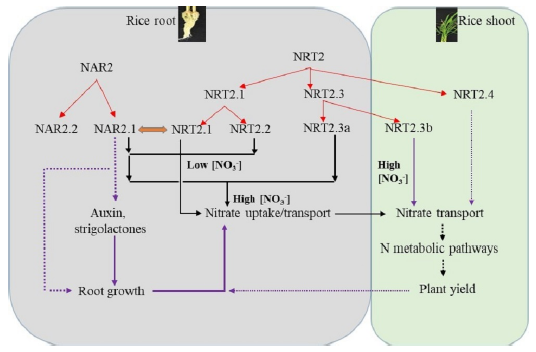
Fig. 2. Schematic representation of proposed evolution and predicted functions for nitrate transporters (NAR2/NRT2) in rice.NAR2.1, NAR2.2, NRT2.1, NRT2.2 and NRT2.3a are expressed mainly in roots; NRT2.3b and NRT2.4 are expressed mainly in shoots (Feng et al, 2011; Yan et al, 2011). Both NRT2.1 and NRT2.2 associated with the nitrate transport (NAR2.1) in the high-affinity concentration range. NRT2.3a requires NAR2.1 for the nitrate transport function, and the protein has a 10-fold lower affinity for nitrate than NRT2.1 and NRT2.2. NAR2.1 can provide a switch, depending on the partner transporter, to enable a rapid response in nitrate uptake over the dynamic ranges of external concentrations of nitrate (Feng et al, 2011; Yan et al, 2011). In contrast, NRT2.3b can function in nitrate transport independently, mainly in the shoot, and its overexpression can greatly improve nitrogen use efficiency and grain yield in rice (Feng et al, 2011; Yan et al, 2011). The solid red arrows represent defined direct functions of the transporters in nitrate uptake and translocation; the dashed arrows represent presumed relationships based on the tissue localization of the genes in rice plant and functional expression in oocytes. The blue arrows indicate the proposed evolution of individual members of the NAR2 and NRT2 nitrate transporter families. Black arrows indicate the possible relationships between NAR2.1 and root growth and between the functions of NRT2 members and plant growth and development.
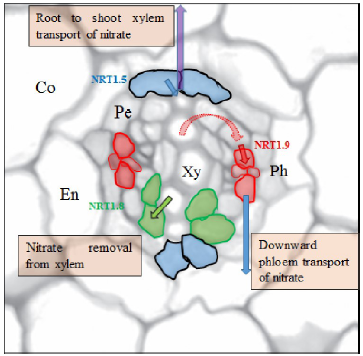
Fig. 3. Regulation of root-to-shoot transport of nitrate.Nitrate transporter 1.5 (NRT1.5), expressed in the xylem pole pericycle, mediates xylem-loading of nitrate (red arrow) and facilitates root-to-shoot xylem transport of nitrate (blue arrow). NRT1.9, expressed in companion cells of phloem, mediates phloem loading (dark-purple arrow) for downward phloem transport of nitrate (dark blue arrow), which may facilitate xylem-to-phloem transfer of nitrate (broken purple arrow), and then reduce root-to-shoot nitrate transport. NRT1.8, expressed in xylem parenchyma, mediates nitrate removal from xylem (green arrow), and could also reduce root-to-shoot nitrate transport.Co, Cortex; En, Endodermis; Pe, Pericycle; Ph, Phloem; Xy, Xylem.
| [1] | Bellegarde F, Gojon A, Martin A.2017. Signals and players in the transcriptional regulation of root responses by local and systemic N signaling in Arabidopsis thaliana. J Exp Bot, 68: 2553-2565. |
| [2] | Bloom A J, Rubio-Asensio J S.2017. Inorganic nitrogen form: A major player in wheat and Arabidopsis responses to elevated CO2. J Exp Bot, 68: 2611-2625. |
| [3] | Bouguyon E, Brun F, Meynard D, Kubeš M, Pervent M, Leran S, Lacombe B, Krouk G, Guiderdoni E, Zažímalová E, Hoyerová K, Nacry P, Gojon A.2015. Multiple mechanisms of nitrate sensing by Arabidopsis nitrate transceptor NRT1.1. Nat Plants, 1: 15015. |
| [4] | Bouguyon E, Perrine-Walker F, Pervent M, Rochette J, Cuesta C, Benkova E, Martiniere A, Bach L, Krouk G, Gojon A, Nacry P.2016. Nitrate controls root development through post- transcriptional regulation of the NRT1.1/NPF6.3 transporter/ sensor.Plant Physiol, 172(2): 1237-1248. |
| [5] | Cai C, Wang J Y, Zhu Y G, Shen Q R, Li B, Tong Y P, Li Z S.2008. Gene structure and expression of the high-affinity nitrate transport system in rice roots.J Integr Plant Biol, 50(4): 443-451. |
| [6] | Castaings L, Camargo A, Pocholle D, Gaudon V, Texier Y, Boutet-Mercey S, Taconnat L, Renou J P, Daniel-Vedele F, Fernandez E, Meyer C, Krapp A.2009. The nodule inception- like protein 7 modulates nitrate sensing and metabolism inArabidopsis. Plant J, 57(3): 426-435. |
| [7] | Chen X B, Yao Q F, Gao X H, Jiang C F, Harberd N P, Fu X D.2016. Shoot-to-root mobile transcription factor HY5 coordinates plant carbon and nitrogen acquisition.Curr Biol, 26(5): 640-646. |
| [8] | Correa L G G, Riano-Pachon D M, Schrago C G, dos Santos R V, Mueller-Roeber B, Vincentz M.2008. The role of bZIP transcription factors in green plant evolution: Adaptive features emerging from four founder genes.PLoS One, 3(8): e2944. |
| [9] | Coruzzi G M, Zhou L.2001. Carbon and nitrogen sensing and signaling in plants: Emerging ‘matrix effects’.Curr Opin Plant Biol, 4(3): 247-253. |
| [10] | Dechorgnat J, Nguyen C T, Armengaud P, Jossier M, Diatloff E, Filleur S, Daniel-Vedele F.2011. From the soil to the seeds: The long journey of nitrate in plants.J Exp Bot, 62(4): 1349-1359. |
| [11] | Espen L, Nocito F F, Cocucci M.2004. Effect of NO3-transport and reduction on intracellular pH: An in vivo NMR study in maize roots. J Exp Bot, 55: 2053-2061. |
| [12] | Fan X R, Feng H M, Tan Y W, Xu Y L, Miao Q S, Xu G H.2016a. A putative 6-transmembrane nitrate transporter OsNRT1.1b plays a key role in rice under low nitrogen.J Integr Plant Biol, 58(6): 590-599. |
| [13] | Fan X R, Tang Z, Tan Y W, Zhang Y, Luo B B, Yang M, Lian X M, Shen Q R, Miller A J, Xu G H.2016b. Overexpression of a pH-sensitive nitrate transporter in rice increases crop yields.Proc Natl Acad Sci USA, 113: 7118-7123. |
| [14] | Feng H M, Yan M, Fan X R, Li B Z, Shen Q R, Miller A J, Xu G H.2011. Spatial expression and regulation of rice high-affinity nitrate transporters by nitrogen and carbon status.J Exp Bot, 62(7): 2319-2332. |
| [15] | Ferreira L M, de Souza V M, Tavares O C H, Zonta E, Santa- Catarina C, de Souza S R, Fernandes M S, Santos L A.2015. OsAMT1.3 expression alters rice ammonium uptake kinetics and root morphology. Plant Biotechnol Rep, 9(4): 221-229. |
| [16] | Forde B G.2002. Local and long-range signaling pathways regulating plant responses to nitrate.Annu Rev Plant Biol, 53: 203-224. |
| [17] | Gojon A, Nacry P, Davidian J C.2009. Root uptake regulation: A central process for NPS homeostasis in plants.Curr Opin Plant Biol, 12(3): 328-338. |
| [18] | Gojon A, Krouk G, Perrine-Walker F, Laugier E.2011. Nitrate transceptor(s) in plants.J Exp Bot, 62(7): 2299-2308. |
| [19] | Gu C S, Song A P, Zhang X X, Wang H B, Li T, Chen Y, Jiang J F, Chen F D, Chen S M.2016. Cloning of chrysanthemum high affinity nitrate transporter family (CmNRT2) and characterization of CmNRT2.1.Sci Rep, 6: 23462. |
| [20] | Guan P Z, Wang R C, Nacry P, Breton G, Kay S A, Pruneda-Paz J L, Davani A, Crawford N M.2014. Nitrate foraging by Arabidopsis roots is mediated by the transcription factor TCP20 through the systemic signaling pathway. Proc Natl Acad Sci USA, 111: 15267-15272. |
| [21] | He M Z, Dijkstra F A.2014. Drought effect on plant nitrogen and phosphorus: A meta analysis.New Phytol, 204(4): 924-931. |
| [22] | Ho C H, Lin S H, Hu H C, Tsay Y F.2009. CHL1 functions as a nitrate sensor in plants.Cell, 138(6): 1184-1194. |
| [23] | Ho C H, Tsay Y F.2010. Nitrate, ammonium, and potassium sensing and signaling.Curr Opin Plant Biol, 13(5): 604-610. |
| [24] | Hoffland E, Dicke M, van Tintelen W, Dijkman H, van Beusichem M L.2000. Nitrogen availability and defense of tomato against two-spotted spider mite.J Chem Ecol, 26(12): 2697-2711. |
| [25] | Holsbeeks I, Lagatie O, van Nuland A, van de Velde S, Thevelein J M.2004. The eukaryotic plasma membrane as a nutrient- sensing device.Trends Biochem Sci, 29(10): 556-564. |
| [26] | Hu B, Wang W, Ou S J, Tang J Y, Li H, Che R H, Zhang Z H, Chai X Y, Wang H R, Wang Y Q, Liang C Z, Liu L C, Piao Z Z, Deng Q Y, Deng K, Xu C, Liang Y, Zhang L H, Li L G, Chu C C.2015. Variation in NRT1.1B contributes to nitrate-use divergence between rice subspecies.Nat Genet, 47: 834-838. |
| [27] | Huang N C, Liu K H, Lo H J, Tsay Y F.1999. Cloning and functional characterization of an Arabidopsis nitrate transporter gene that encodes a constitutive component of low-affinity uptake. Plant Cell, 11(8): 1381-1392. |
| [28] | Jakoby M, Weisshaar B, Droge-Laser W, Vicente-Carbajosa J, Tiedemann J, Kroj T, Parcy F.2002. bZIP transcription factors in Arabidopsis. Trends Plant Sci, 7(3): 106-111. |
| [29] | Jonassen E M, Lea U S, Lillo C.2008. HY5 and HYH are positive regulators of nitrate reductase in seedlings and rosette stage plants.Planta, 227(3): 559-564. |
| [30] | Kanno Y, Hanada A, Chiba Y, Ichikawa T, Nakazawa M, Matsui M, Koshiba T, Kamiya Y, Seo M.2012. Identification of an abscisic acid transporter by functional screening using the receptor complex as a sensor.Proc Natl Acad Sci USA, 109: 9653-9658. |
| [31] | Kawachi T, Sunaga Y, Ebato M, Hatanaka T, Harada H.2006. Repression of nitrate uptake by replacement of Asp105 by asparagine in AtNRT3.1 in Arabidopsis thaliana L. Plant Cell Physiol, 47(10): 1437-1441. |
| [32] | Kiba T, Feria-Bourrellier A B, Lafouge F, Lezhneva L, Boutet-Mercey S, Orsel M, Bréhaut V, Miller A, Daniel-Vedele F, Sakakibara H, Krapp A.2012. The Arabidopsis nitrate transporter NRT2.4 plays a double role in roots and shoots of nitrogen-starved plants. Plant Cell, 24(1): 245-258. |
| [33] | Kiba T, Krapp A.2016. Plant nitrogen acquisition under low availability: Regulation of uptake and root architecture.Plant Cell Physiol, 57(4): 707-714. |
| [34] | Kotur Z, Mackenzie N, Ramesh S, Tyerman S D, Kaiser B N, Glass A D.2012. Nitrate transport capacity of the Arabidopsis thaliana NRT2 family members and their interactions with AtNAR2.1. New Phytol, 194(3): 724-731. |
| [35] | Kotur Z, Glass A D.2015. A 150 kDa plasma membrane complex of AtNRT2.5 and AtNAR2.1 is the major contributor to constitutive high affinity nitrate influx in Arabidopsis thaliana. Plant Cell Environ, 38(8): 1490-1502. |
| [36] | Krapp A, David L C, Chardin C, Girin T, Marmagne A, Leprince A S, Chaillou S, Ferrario-Mery S, Meyer C, Daniel-Vedele F.2014. Nitrate transport and signalling inArabidopsis. J Exp Bot, 65(3): 789-798. |
| [37] | Krouk G, Tillard P, Gojon A.2006. Regulation of the high-affinity NO3- uptake system by NRT1.1-mediated NO3- demand signaling inArabidopsis. Plant Physiol, 142(3): 1075-1086. |
| [38] | Krouk G, Lacombe B, Bielach A, Perrine-Walker F, Malinska K, Mounier E, Hoyerova K, Tillard P, Leon S, Ljung K, Zazimalova E, Benkova E, Nacry P, Gojon A.2010. Nitrate- regulated auxin transport by NRT1.1 defines a mechanism for nutrient sensing in plants.Dev Cell, 18(6): 927-937. |
| [39] | Lanquar V, Loque D, Hormann F, Yuan L X, Bohner A, Engelsberger W R, Lalonde S, Schulze W X, von Wiren N, Frommer W B.2009. Feedback inhibition of ammonium uptake by a phospho-dependent allosteric mechanism in Arabidopsis. Plant Cell, 21(11): 3610-3622. |
| [40] | Laugier E, Bouguyon E, Mauriès A, Tillard P, Gojon A, Lejay L.2012. Regulation of high-affinity nitrate uptake in roots of Arabidopsis depends predominantly on posttranscriptional control of the NRT2.1/NAR2.1 transport system. Plant Physiol, 158(2): 1067-1078. |
| [41] | le Deunff E, Lecourt J, Malagoli P.2016. Fine-tuning of root elongation by ethylene a tool to study dynamic structure- function relationships between root architecture and nitrate absorption.Ann Bot, 118(4): 607-620. |
| [42] | Léran S, Muños S, Brachet C, Tillard P, Gojon A, Lacombe B.2013. Arabidopsis NRT1.1 is a bidirectional transporter involved in root-to-shoot nitrate translocation. Mol Plant, 6: 1984-1987. |
| [43] | Lezhneva L, Kiba T, Feria-Bourrellier A B, Lafouge F, Boutet- Mercey S, Zoufan P, Sakakibara H, Daniel-Vedele F, Krapp A.2014. The Arabidopsis nitrate transporter NRT2.5 plays a role in nitrate acquisition and remobilization in nitrogen-starved plants. Plant J, 80(2): 230-241. |
| [44] | Li C, Tang Z, Wei J, Qu H Y, Xie Y J, Xu G H.2016. The OsAMT1.1 gene functions in ammonium uptake and ammonium- potassium homeostasis over low and high ammonium concentration ranges. J Genet Genom, 43(11): 639-649. |
| [45] | Li J Y, Fu Y L, Pike S M, Bao J, Tian W, Zhang Y, Chen C Z, Zhang Y, Li H M, Huang J, Li L G, Schroeder J I, Gassmann W, Gong J M.2010. The Arabidopsis nitrate transporter NRT1.8 functions in nitrate removal from the xylem sap and mediates cadmium tolerance. Plant Cell, 22(5): 1633-1646. |
| [46] | Li W B, Wang Y, Okamoto M, Crawford N M, Siddiqi M Y, Glass A D M.2007. Dissection of the AtNRT2.1:AtNRT2.2 inducible high-affinity nitrate transporter gene cluster.Plant Physiol, 143(1): 425-433. |
| [47] | Li Y F, Zheng Y, Addo-Quaye C, Zhang L, Saini A, Jagadeeswaran G, Axtell M J, Zhang W X, Sunkar R.2010. Transcriptome-wide identification of microRNA targets in rice.Plant J, 62(5): 742-759. |
| [48] | Lima J E, Kojima S, Takahashi H, von Wirén N.2010. Ammonium triggers lateral root branching in Arabidopsis in an AMMONIUM TRANSPORTER1;3-dependent manner. Plant Cell, 22(11): 3621-3633. |
| [49] | Lin S H, Kuo H F, Canivenc G, Lin C S, Lepetit M, Hsu P K, Tillard P, Lin H L, Wang Y Y, Tsai C B, Gojon A, Tsay Y F.2008. Mutation of the Arabidopsis NRT1.5 nitrate transporter causes defective root-to-shoot nitrate transport. Plant Cell, 20(9): 2514-2528. |
| [50] | Lin Y L, Tsay Y F.2017. Influence of differing nitrate and nitrogen availability on flowering control in Arabidopsis. J Exp Bot, 68(10): 2603-2609. |
| [51] | Little D Y, Rao H, Oliva S, Daniel-Vedele F, Krapp A, Malamy J E.2005. The putative high-affinity nitrate transporter NRT2.1 represses lateral root initiation in response to nutritional cues.Proc Natl Acad Sci USA, 102: 13693-13698. |
| [52] | Liu K H, Tsay Y F.2003. Switching between the two action modes of the dual-affinity nitrate transporter CHL1 by phosphorylation.EMBO J, 22(5): 1005-1013. |
| [53] | Liu W W, Sun Q, Wang K, Du Q G, Li W X.2017. Nitrogen limitation adaptation (NLA) is involved in source-to-sink remobilization of nitrate by mediating the degradation of NRT1.7 in Arabidopsis. New Phytol, 214(2): 734-744. |
| [54] | Liu X Q, Huang D M, Tao J Y, Miller A J, Fan X R, Xu G H.2014. Identification and functional assay of the interaction motifs in the partner protein OsNAR2.1 of the two-component system for high-affinity nitrate transport.New Phytol, 204(1): 74-80. |
| [55] | Liu Y, von Wirén N.2017. Ammonium as a signal for physiological and morphological responses in plants.J Exp Bot, 68(10): 2581-2592. |
| [56] | Malamy J E.2005. Intrinsic and environmental response pathways that regulate root system architecture.Plant Cell Environ, 28(1): 67-77. |
| [57] | Marchive C, Roudier F, Castaings L, Bréhaut V, Blondet E, Colot V, Meyer C, Krapp A.2013. Nuclear retention of the transcription factor NLP7 orchestrates the early response to nitrate in plants.Nat Commun, 4: 1713. |
| [58] | Moran-Zuloaga D, Dippold M, Glaser B, Kuzyakov Y.2015. Organic nitrogen uptake by plants: Reevaluation by position- specific labeling of amino acids.Biogeochem, 125(3): 359-374. |
| [59] | Morère-Le Paven M C, Viau L, Hamon A, Vandecasteele C, Pellizzaro A, Bourdin C, Laffont C, Lapied B, Lepetit M, Frugier F, Legros C, Limami A M.2011. Characterization of a dual-affinity nitrate transporter MtNRT1.3 in the model legume Medicago truncatula. J Exp Bot, 62: 5595-5605. |
| [60] | Mounier E, Pervent M, Ljung K, Gojon A, Nacry P.2014. Auxinmediated nitrate signalling by NRT1.1 participates in the adaptive response of Arabidopsis root architecture to the spatial heterogeneity of nitrate availability. Plant Cell Environ, 37(1): 162-174. |
| [61] | Munõs S, Cazettes C, Fizames C, Gaymard F, Tillard P, Lepetit M, Lejay L, Gojon A.2004. Transcript profiling in the chl1-5 mutant of Arabidopsis reveals a role of the nitrate transporter NRT1.1 in the regulation of another nitrate transporter, NRT2.1. Plant Cell, 16(9): 2433-2447. |
| [62] | Näsholm T, Kielland K, Ganeteg U.2009. Uptake of organic nitrogen by plants.New Phytol, 182(1): 31-48. |
| [63] | Nijhawan A, Jain M, Tyagi A K, Khurana J P.2008. Genomic survey and gene expression analysis of the basic leucine zipper transcription factor family in rice.Plant Physiol, 146(2): 333-350. |
| [64] | O’Brien J A, Vega A, Bouguyon E, Krouk G, Gojon A, Coruzzi G, Gutiérrez R A.2016. Nitrate transport, sensing, and responses in plants.Mol Plant, 9(6): 837-856. |
| [65] | Ohkubo Y, Tanaka M, Tabata R, Ogawa-Ohnishi M, Matsubayashi Y.2017. Shoot-to-root mobile polypeptides involved in systemic regulation of nitrogen acquisition.Nat Plants, 3: 17029. |
| [66] | Okamoto M, Kumar A, Li W B, Wang Y, Siddiqi M Y, Crawford N M, Glass A D.2006. High-affinity nitrate transport in roots of Arabidopsis depends on expression of the NAR2-like gene AtNRT3.1. Plant Physiol, 140(3): 1036-1046. |
| [67] | Okamoto S, Shinohara H, Mori T, Matsubayashi Y, Kawaguchi M.2013. Root-derived CLE glycopeptides control nodulation by direct binding to HAR1 receptor kinase.Nat Commun, 4: 2191. |
| [68] | Okamoto S, Tabata R, Matsubayashi Y.2016. Long-distance peptide signaling essential for nutrient homeostasis in plants.Curr Opin Plant Biol, 34: 35-40. |
| [69] | Orsel M, Chopin F, Leleu O, Smith S J, Krapp A, Daniel-Vedele F, Miller A J.2006. Characterization of a two-component high- affinity nitrate uptake system in Arabidopsis: Physiology and protein-protein interaction. Plant Physiol, 142(3): 1304-1317. |
| [70] | Pacheco-Villalobos D, Diaz-Moreno S M, van der Schuren A, Tamaki T, Kang Y H, Gujas B, Novak O, Jaspert N, Li Z, Wolf S, Oecking C, Ljung K, Bulone V, Hardtke C S.2016. The effects of high steady state auxin levels on root cell elongation in Brachypodium. Plant Cell, 28(5): 1009-1024. |
| [71] | Parker J L, Newstead S.2014. Molecular basis of nitrate uptake by the plant nitrate transporter NRT1.1.Nature, 507: 68-72. |
| [72] | Plett D, Toubia J, Garnett T, Tester M, Kaiser B N, Baumann U.2010. Dichotomy in the NRT gene families of dicots and grass species. PLoS One, 5: e15289. |
| [73] | Rahayu Y S, Walch-Liu P, Neumann G, Romheld V, von Wiren N, Bangerth F.2005. Root-derived cytokinins as long-distance signals for NO3- induced stimulation of leaf growth.J Exp Bot, 56: 1143-1152. |
| [74] | Ranathunge K, El-kereamy A, Gidda S, Bi Y M, Rothstein S J.2014. AMT1;1 transgenic rice plants with enhanced NH4 permeability show superior growth and higher yield under optimal and suboptimal NH4 conditions.J Exp Bot, 65(4): 965-979. |
| [75] | Reguera M, Peleg Z, Abdel-Tawab Y M, Tumimbang E B, Delatorre C A, Blumwald E.2013. Stress-induced cytokinin synthesis increases drought tolerance through the coordinated regulation of carbon and nitrogen assimilation in rice.Plant Physiol, 163(4): 1609-1622. |
| [76] | Remans T, Nacry P, Pervent M, Filleur S, Diatloff E, Mounier E, Tillard P, Forde B G, Gojon A.2006. The Arabidopsis NRT1.1 transporter participates in the signaling pathway triggering root colonization of nitraterich patches. Proc Natl Acad Sci USA, 103: 19206-19211. |
| [77] | Riveras E, Alvarez J M, Vidal E A, Oses C, Vega A, Gutiérrez R A.2015. The calcium ion is a second messenger in the nitrate signaling pathway of Arabidopsis. Plant Physiol, 169(2): 1397-1404. |
| [78] | Rubio-Covarrubias O A, Brown P H, Weinbaum S A, Johnson R S, Cabrera R I.2009. Evaluating foliar nitrogen compounds as indicators of nitrogen status in Prunus persica trees. Sci Hort, 120(1): 27-33. |
| [79] | Ruffel S, Krouk G, Ristova D, Shasha D, Birnbaum K D, Coruzzi G M.2011. Nitrogen economics of root foraging: Transitive closure of the nitrate-cytokinin relay and distinct systemic signaling for N supply vs. demand.Proc Natl Acad Sci USA, 108: 18524-18529. |
| [80] | Schachtman D P, Shin R.2007. Nutrient sensing and signaling: NPKS.Annu Rev Plant Biol, 58: 47-69. |
| [81] | Segonzac C, Boyer J C, Ipotesi E, Szponarski W, Tillard P, Touraine B, Sommerer N, Rossignol M, Gibrat R.2007. Nitrate efflux at the root plasma membrane: Identification of an Arabidopsis excretion transporter. Plant Cell, 19(11): 3760-3777. |
| [82] | Sinfield J V, Fagerman D, Colic O.2010. Evaluation of sensing technologies for on-the-go detection of macro-nutrients in cultivated soils.Comput Electron Agric, 70(1): 1-18. |
| [83] | Sonoda Y, Ikeda A, Saiki S, von Wiren N, Yamaya T, Yamaguchi J.2003a. Distinct expression and function of three ammonium transporter genes (OsAMT1;1-1;3) in rice. Plant Cell Physiol, 44(7): 726-734. |
| [84] | Sonoda Y, Ikeda A, Saiki S, Yamaya T, Yamaguchi J.2003b. Feedback regulation of the ammonium transporter gene family AMT1 by glutamine in rice.Plant Cell Physiol, 44(12): 1396-1402. |
| [85] | Stitt M.1999. Nitrate regulation of metabolism and growth.Curr Opin Plant Biol, 2: 178-186. |
| [86] | Straub T, Ludewig U, Neuhaeuser B.2017. The kinase CIPK23 inhibits ammonium transport in Arabidopsis thaliana. Plant Cell, 29(2): 409-422. |
| [87] | Suenaga A, Moriya K, Sonoda Y, Ikeda A, von Wiren N, Hayakawa T, Yamaguchi J, Yamaya T.2003. Constitutive expression of a novel-type ammonium transporter OsAMT2 in rice plants.Plant Cell Physiol, 44(2): 206-211. |
| [88] | Sui Z P, Wang T Y, Li H J, Zhang M, Li Y Y, Xu R B, Xing G F, Ni Z F, Xin M M.2016. Overexpression of peptide-encoding OsCEP6.1 results in pleiotropic effects on growth in rice (O. sativa). Front Plant Sci, 7: 228. |
| [89] | Sun J, Bankston J R, Payandeh J, Hinds T R, Zagotta W N, Zheng N.2014. Crystal structure of the plant dual-affinity nitrate transporter NRT1.1.Nature, 507: 73-77. |
| [90] | Tabata R, Sumida K, Yoshii T, Ohyama K, Shinohara H, Matsubayashi Y.2014. Perception of root-derived peptides by shoot LRR-RKs mediates systemic N-demand signaling.Science, 346: 343-346. |
| [91] | Taiz L, Zeiger E.2010. Plant Physiology. 5th edn. Sunderland, MA, USA: Sinauer Associates Inc: 67-86. |
| [92] | Tang Z, Fan X R, Li Q, Feng H M, Miller A J, Shen Q R, Xu G H.2012. Knockdown of a rice stelar nitrate transporter alters long-distance translocation but not root influx.Plant Physiol, 160: 2052-2063. |
| [93] | Taochy C, Gaillard I, Ipotesi E, Oomen R, Leonhardt N, Zimmermann S, Peltier J B, Szponarski W, Simonneau T, Sentenac H, Gibrat R, Boyer J C.2015. The Arabidopsis root stele transporter NPF2.3 contributes to nitrate translocation to shoots under salt stress.Plant J, 83(3): 466-479. |
| [94] | Taulemesse F, le Gouis J, Gouache D, Gibon Y, Allard V.2015. Post flowering nitrate uptake in wheat is controlled by N status at flowering, with a putative major role of root nitrate transporter NRT2.1.PLoS One, 10(3): e0120291. |
| [95] | Tremblay N, Fallon E, Ziadi N.2011. Sensing of crop nitrogen status: Opportunities, tools, limitations, and supporting information requirements.Hort Technol, 21: 274-281. |
| [96] | Tsay Y F, Schroeder J I, Feldmann K A, Crawford N M.1993. The herbicide sensitivity gene CHL1 of Arabidopsis encodes a nitrate-inducible nitrate transporter. Cell, 72(5): 705-713. |
| [97] | Tsay Y F, Chiu C C, Tsai C B, Ho C H, Hsu P K.2007. Nitrate transporters and peptide transporters.FEBS Lett, 581: 2290-2300. |
| [98] | Tsay Y F.2014. Plant science: How to switch affinity.Nature, 507: 44-45. |
| [99] | Undurraga S F, Ibarra-Henríquez C, Fredes I, Miguel-Álvarez J M, Gutiérrez R A.2017. Nitrate signaling and early responses in Arabidopsis roots. J Exp Bot, 68(10): 2541-2551. |
| [100] | Verbon E H, Liberman L M.2016. Beneficial microbes affect endogenous mechanisms controlling root development.Trends Plant Sci, 2(3): 218-229. |
| [101] | Vidal E A, Araus V, Lu C, Parry G, Green P J, Coruzzi G M, Gutierrez R A.2010. Nitrate-responsive miR393/AFB3 regulatory module controls root system architecture in Arabidopsis thaliana. Proc Natl Acad Sci USA, 107(9): 4477-4482. |
| [102] | Vidal E A, Moyano T C, Krouk G, Katari M S, Tanurdzic M, McCombie W R, Coruzzi G M, Gutierrez R A.2013. Integrated RNA-seq and sRNA-seq analysis identifies novel nitrate- responsive genes inArabidopsis thaliana roots. BMC Genom, 14: 701. |
| [103] | Vigneau N, Ecarnot M, Rabatel G, Roumet P.2011. Potential of field hyperspectral imaging as a non-destructive method to assess leaf nitrogen content in wheat.Field Crops Res, 122(1): 25-31. |
| [104] | Wang R C, Xing X J, Wang Y, Tran A, Crawford N M.2009. A genetic screen for nitrate regulatory mutants captures the nitrate transporter geneNRT1.1. Plant Physiol, 151(1): 472-478. |
| [105] | Wang W, Hu B, Yuan D Y, Liu Y Q, Che R H, Hu Y C, Ou S J, Liu Y X, Zhang Z H, Wang H R, Li H, Jiang Z M, Zhang Z L, Gao X K, Qiu Y H, Meng X B, Liu Y X, Bai Y, Liang Y, Wang Y Q, Zhang L H, Li L G, Sodmergen, Jing H C, Li J Y, Chu C C.2018. Expression of the nitrate transporter gene OsNRT1.1A/ OsNPF6.3 confers high yield and early maturation in rice. Plant Cell, 30(3): 638-651. |
| [106] | Wang Y Y, Hsu P K, Tsay Y F.2012. Uptake, allocation and signaling of nitrate.Trends Plant Sci, 17(8): 458-467. |
| [107] | Wen Z Y, Tyerman S D, Dechorgnat J, Ovchinnikova E, Dhugga K S, Kaiser B N.2017. Maize NPF6 proteins are homologs of Arabidopsis CHL1 that are selective for both nitrate and chloride. Plant Cell, 29(10): 2581-2596. |
| [108] | Xia X D, Fan X R, Wei J, Feng H M, Qu H Y, Xie D, Miller A J, Xu G H.2015. Rice nitrate transporter OsNPF2.4 functions in low-affinity acquisition and long-distance transport.J Exp Bot, 66(1): 317-331. |
| [109] | Xu G H, Fan X R, Miller A J.2012. Plant nitrogen assimilation and use efficiency.Annu Rev Plant Biol, 63: 153-182. |
| [110] | Xuan W, Band L R, Kumpf R P, van Damme D, Parizot B, de Rop G, Opdenacker D, Moller B K, Skorzinski N, Njo M F, de Rybel B, Audenaert D, Nowack M K, Vanneste S, Beeckman T.2016. Cyclic programmed cell death stimulates hormone signaling and root development in Arabidopsis. Science, 351: 384-387. |
| [111] | Yan M, Fan X R, Feng H M, Miller A J, Shen Q R, Xu G H.2011. Rice OsNAR2.1 interacts with OsNRT2.1, OsNRT2.2 and OsNRT2.3a nitrate transporters to provide uptake over high and low concentration ranges.Plant Cell Environ, 34(8): 1360-1372. |
| [112] | Yan Y S, Wang H C, Hamera S, Chen X Y, Fang R X.2014. miR444a has multiple functions in the rice nitrate-signaling pathway.Plant J, 78(1): 44-55. |
| [113] | Yang S Y, Hao D L, Cong Y, Jin M, Su Y H.2015. The rice OsAMT1;1 is a proton-independent feedback regulated ammonium transporter.Plant Cell Rep, 34(2): 321-330. |
| [114] | Yang W Z, Yoon J, Choi H, Fan Y L, Chen R M, An G.2015. Transcriptome analysis of nitrogen-starvation-responsive genes in rice.BMC Plant Biol, 15: 31. |
| [115] | Yong Z H, Kotur Z, Glass A D M.2010. Characterization of an intact two component high-affinity nitrate transporter from Arabidopsis roots. Plant J, 63(5): 739-748. |
| [116] | Yu C Y, Su S, Xu Y C, Zhao Y Q, Yan A, Huang L L, Ali I, Gan Y B.2014. The effects of fluctuations in the nutrient supply on the expression of five members of the AGL17 clade of MADS-box genes in rice.PLoS One, 9(8): e105597. |
| [117] | Yu C Y, Liu Y H, Zhang A D, Su S, Yan A, Huang L, Ali I, Liu Y, Forde B G, Gan Y B.2015. MADS-box transcription factor OsMADS25 regulates root development through affection of nitrate accumulation in rice.PLoS One, 10(8): e0135196. |
| [118] | Yuan S, Zhang Z W, Zheng C, Zhao Z Y, Wang Y, Feng L Y, Niu G Q, Wang C Q, Wang J H, Feng H, Xu F, Bao F, Hu Y, Cao Y, Ma L G, Wang H Y, Kong D D, Xiao W, Lin H H, He Y K.2016. Arabidopsis cryptochrome 1 functions in nitrogen regulation of flowering. Proc Natl Acad Sci USA, 113: 7661-7666. |
| [119] | Zhang H, Forde B G.1998. An Arabidopsis MADS box gene that controls nutrient-induced changes in root architecture. Science, 279: 407-409. |
| [120] | Zhang X, Davidson E A, Mauzerall D L, Searchinger T D, Dumas P, Shen Y.2015. Managing nitrogen for sustainable development.Nature, 528: 51-59. |
| [121] | Zhao M, Ding H, Zhu J K, Zhang F S, Li W X.2011. Involvement of miR169 in the nitrogen-starvation responses in Arabidopsis. New Phytol, 190(4): 906-915. |
| [122] | Zhu Y Y, Di T J, Xu G H, Chen X, Zeng H Q, Yan F, Shen Q R.2009. Adaptation of plasma membrane H+-ATPase of rice roots to low pH as related to ammonium nutrition.Plant Cell Environ, 32: 1428-1440. |
| [1] | Prathap V, Suresh KUMAR, Nand Lal MEENA, Chirag MAHESHWARI, Monika DALAL, Aruna TYAGI. Phosphorus Starvation Tolerance in Rice Through a Combined Physiological, Biochemical and Proteome Analysis [J]. Rice Science, 2023, 30(6): 8-. |
| [2] | Serena REGGI, Elisabetta ONELLI, Alessandra MOSCATELLI, Nadia STROPPA, Matteo Dell’ANNO, Kiril PERFANOV, Luciana ROSSI. Seed-Specific Expression of Apolipoprotein A-IMilano Dimer in Rice Engineered Lines [J]. Rice Science, 2023, 30(6): 6-. |
| [3] | Sundus ZAFAR, XU Jianlong. Recent Advances to Enhance Nutritional Quality of Rice [J]. Rice Science, 2023, 30(6): 4-. |
| [4] | Kankunlanach KHAMPUANG, Nanthana CHAIWONG, Atilla YAZICI, Baris DEMIRER, Ismail CAKMAK, Chanakan PROM-U-THAI. Effect of Sulfur Fertilization on Productivity and Grain Zinc Yield of Rice Grown under Low and Adequate Soil Zinc Applications [J]. Rice Science, 2023, 30(6): 9-. |
| [5] | FAN Fengfeng, CAI Meng, LUO Xiong, LIU Manman, YUAN Huanran, CHENG Mingxing, Ayaz AHMAD, LI Nengwu, LI Shaoqing. Novel QTLs from Wild Rice Oryza longistaminata Confer Rice Strong Tolerance to High Temperature at Seedling Stage [J]. Rice Science, 2023, 30(6): 14-. |
| [6] | LIN Shaodan, YAO Yue, LI Jiayi, LI Xiaobin, MA Jie, WENG Haiyong, CHENG Zuxin, YE Dapeng. Application of UAV-Based Imaging and Deep Learning in Assessment of Rice Blast Resistance [J]. Rice Science, 2023, 30(6): 10-. |
| [7] | Md. Forshed DEWAN, Md. AHIDUZZAMAN, Md. Nahidul ISLAM, Habibul Bari SHOZIB. Potential Benefits of Bioactive Compounds of Traditional Rice Grown in South and South-East Asia: A Review [J]. Rice Science, 2023, 30(6): 5-. |
| [8] | Raja CHAKRABORTY, Pratap KALITA, Saikat SEN. Phenolic Profile, Antioxidant, Antihyperlipidemic and Cardiac Risk Preventive Effect of Chakhao Poireiton (A Pigmented Black Rice) in High-Fat High-Sugar induced Rats [J]. Rice Science, 2023, 30(6): 11-. |
| [9] | LI Qianlong, FENG Qi, WANG Heqin, KANG Yunhai, ZHANG Conghe, DU Ming, ZHANG Yunhu, WANG Hui, CHEN Jinjie, HAN Bin, FANG Yu, WANG Ahong. Genome-Wide Dissection of Quan 9311A Breeding Process and Application Advantages [J]. Rice Science, 2023, 30(6): 7-. |
| [10] | JI Dongling, XIAO Wenhui, SUN Zhiwei, LIU Lijun, GU Junfei, ZHANG Hao, Tom Matthew HARRISON, LIU Ke, WANG Zhiqin, WANG Weilu, YANG Jianchang. Translocation and Distribution of Carbon-Nitrogen in Relation to Rice Yield and Grain Quality as Affected by High Temperature at Early Panicle Initiation Stage [J]. Rice Science, 2023, 30(6): 12-. |
| [11] | Nazaratul Ashifa Abdullah Salim, Norlida Mat Daud, Julieta Griboff, Abdul Rahim Harun. Elemental Assessments in Paddy Soil for Geographical Traceability of Rice from Peninsular Malaysia [J]. Rice Science, 2023, 30(5): 486-498. |
| [12] | Monica Ruffini Castiglione, Stefania Bottega, Carlo Sorce, Carmelina SpanÒ. Effects of Zinc Oxide Particles with Different Sizes on Root Development in Oryza sativa [J]. Rice Science, 2023, 30(5): 449-458. |
| [13] | Tan Jingyi, Zhang Xiaobo, Shang Huihui, Li Panpan, Wang Zhonghao, Liao Xinwei, Xu Xia, Yang Shihua, Gong Junyi, Wu Jianli. ORYZA SATIVA SPOTTED-LEAF 41 (OsSPL41) Negatively Regulates Plant Immunity in Rice [J]. Rice Science, 2023, 30(5): 426-436. |
| [14] | Ammara Latif, Sun Ying, Pu Cuixia, Noman Ali. Rice Curled Its Leaves Either Adaxially or Abaxially to Combat Drought Stress [J]. Rice Science, 2023, 30(5): 405-416. |
| [15] | Liu Qiao, Qiu Linlin, Hua Yangguang, Li Jing, Pang Bo, Zhai Yufeng, Wang Dekai. LHD3 Encoding a J-Domain Protein Controls Heading Date in Rice [J]. Rice Science, 2023, 30(5): 437-448. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
