
Rice Science ›› 2023, Vol. 30 ›› Issue (1): 36-49.DOI: 10.1016/j.rsci.2022.05.004
• Research Paper • Previous Articles Next Articles
Meng Lu1,2, Tang Mingfeng1,3, Zhu Yuxing1, Tan Longtao1( )
)
Received:2022-01-20
Accepted:2022-05-20
Online:2023-01-28
Published:2022-11-11
Contact:
Tan Longtao
About author:First author contact:This is an open access article under the CC BY-NC-ND license (
Peer review under responsibility of China National Rice Research Institute
Meng Lu, Tang Mingfeng, Zhu Yuxing, Tan Longtao. Knocking-Out OsPDR7 Triggers Up-Regulation of OsZIP9 Expression and Enhances Zinc Accumulation in Rice[J]. Rice Science, 2023, 30(1): 36-49.
Add to citation manager EndNote|Ris|BibTeX
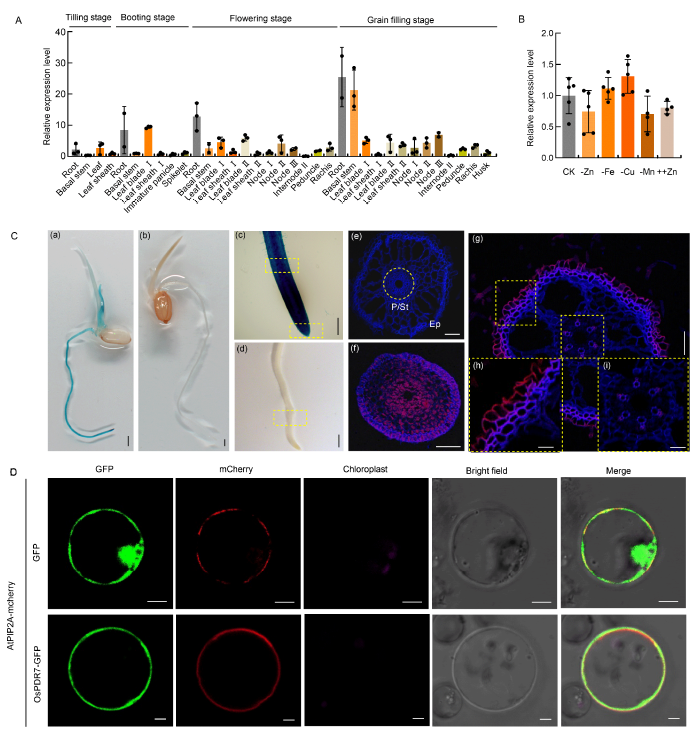
Fig. 1. Expression profile of OsPDR7 and subcellular localization of OsPDR7. A, Tissue specific gene expression in various tissues at different growth stages. Rice plants were grown under field conditions. OsUBC was used as the internal standard for gene expression. I, II and III represent the serial numbers of node, leaf blade, leaf sheath and internode from top to bottom of rice plants. Data are Mean ± SD (n = 3). B, Relative expression levels of OsPDR7 in the roots of seedlings exposed to different metal concentrations. The 21-day-old wild type seedlings were transferred to 0.5× Kimura B (KB) solution deficient in Zn, Fe, Cu and Mn (-Zn, -Fe, -Cu and -Mn) or supplemented with 50 μmol/L Zn (Zn-exposed, ++Zn) for 7 d. Data were separately compared with that from seedlings grown in normal 0.5× KB (CK). Values represent Mean ± SD (n ≥ 3). Statistical comparisons were performed using the Tukey’s HSD mean-separation test at the 0.05 level and no significant difference was observed. C, Histochemical analysis of transgenic plants expressing the GUS gene driven by the OsPDR7 promoter (a and c) and wild type plants (b and d). (a), ProOsPDR7::GUS seedling. (b), Wild type seedling. (c), Primary root of ProOsPDR7::GUS transgenic plant. (d), Primary root of wild type plant. (e-i), Immunostaining with anti-GUS antibody was performed in roots of ProOsPDR7::GUS transgenic rice seedlings (f-i) and wild type plant (e). Blue indicates autofluorescence emitted from the cell wall due to 4,6-diamidino-2-phenylindole (DAPI) staining. Red indicates the anti-GUS antibody-specific fluorescent signal. (e) Representative transverse sections of (d). (f) and (g), Representative transverse sections of (c). (h) and (i), Representative regions magnified from (g). P, Pericycle; St, Stele; Ep, Exodermis. Scale bars, 1 mm in (a-d), 50 μm in (e-g) and 20 μm in (h) and (i). D, Subcellular localization of OsPDR7 protein determined in rice protoplasts. For each localization experiment, ≥ 20 individual cells were examined using a Zeiss LSM880 confocal laser scanning microscope (Carl Zeiss, Germany). Scale bars, 2 μm. GFP, Green fluorescent protein.
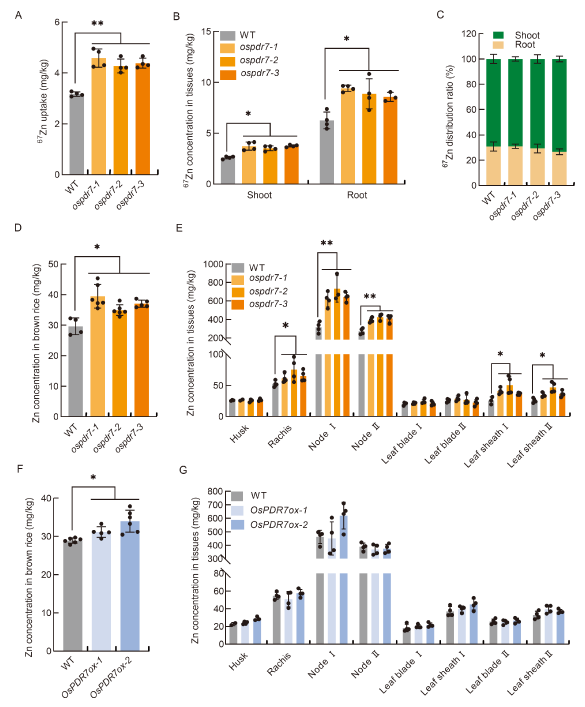
Fig. 2. Zn profiles of OsPDR7 knock-out lines and over- expression lines at vegetative and reproductive growth stages. A, Total uptake of 67Zn by wild type (WT) and three OsPDR7 knock-out lines (ospdr7-1 to -3) at the vegetative stage. B, 67Zn concentration in the shoots and roots at the seedling stage. C, 67Zn distribution ratio between the roots and shoots at the seedling stage. D and E, Zn accumulation in brown rice (D) and in husk, rachis, node, leaf blade and leaf sheath (E) at the reproductive stage. F and G, Zn accumulation in brown rice (F) and in husk, rachis, node, leaf blade and leaf sheath (G) in OsPDR7 over-expression (OsPDR7ox-1 and -2) lines at the reproductive stage. I and II in E and G represent the serial numbers of node, leaf blade and leaf sheath from top to bottom of rice plants.Seedlings were grown in 0.5× Kimura B (KB) nutrient solution for 21 d, then transferred to 0.5× KB containing 0.4 μmol/L 67ZnSO4 for 2 d in A-C. Plants were grown in a paddy field until the grains were ripe in D-G. Values represent Mean ± SD (n ≥ 3). Statistical comparisons against WT were performed with the Tukey’s HSD mean-separation test (*, P < 0.05; **, P < 0.01).
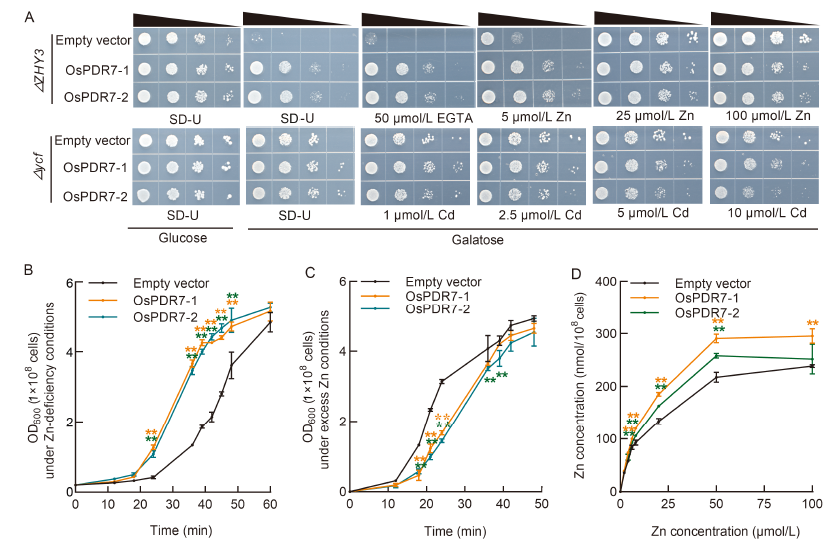
Fig. 3. Complementation assay of OsPDR7 in yeast mutant strain. A, Zn and Cd transport activity assay of OsPDR7 expressed in yeast. The empty vector pYES2C or OsPDR7 was introduced into Zn uptake-deficient mutant strain ?ZHY3 or Cd-sensitive mutant strain ?ycf on the synthetic defined-Ura (SD-U) medium. The yeast strains were cultured on the plate with different Zn or Cd concentrations at 30 for 2-4 d. Zn was chelated with 50 μmol/L ethylenebis (oxyethylenenitrilo) tetraacetic acid (EGTA). B and C, Growth rates of yeast cells. Yeast mutant strain ?ZHY3 expressing OsPDR7 or the empty vector was cultured in SD-U medium containing galactose with 2.5 μmol/L Zn (B) or 500 μmol/L Zn (C). D, Concentration of Zn in the mutant yeast strain ?ZHY3 expressing the empty vector pYES2C or OsPDR7. Data are Mean ± SD (n ≥ 3). In B-D, significant differences were determined by one-way analysis of variance (*, P < 0.05; **, P < 0.01).
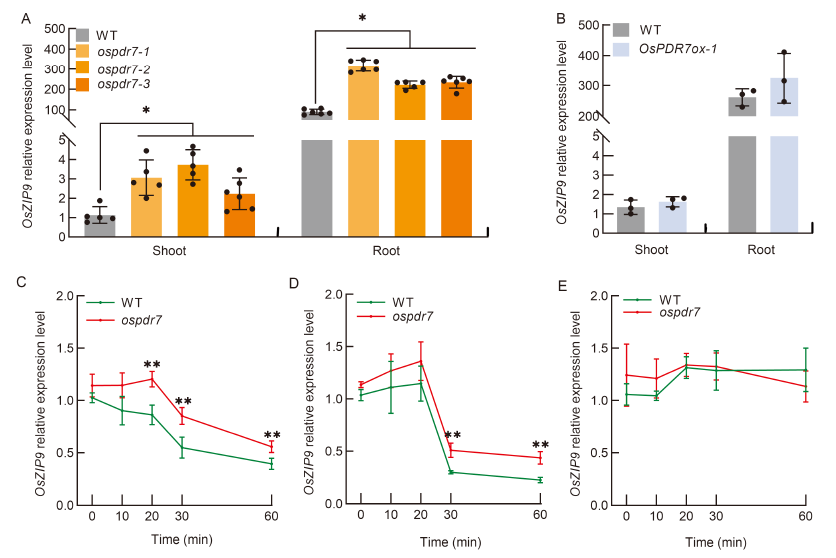
Fig. 4. OsZIP9 expression in response to exogenous Zn supplied via root culture or foliar spray. A and B, Expression levels of OsZIP9 in roots and shoots of wild type (WT) and three OsPDR7 knock-out lines (ospdr7-1 to -3) (A), as well as in WT and OsPDR7 over-expression line (OsPDR7ox-1) (B). The 28-day-old seedlings were used. C-E, OsZIP9 relative expression levels in local system (C), foliar spray with 0.4 μmol/L 67ZnSO4 (D), and in control treatments (E) in WT and OsPDR7 knockout lines. The 21-day-old seedlings were cultivated in Zn-deficient 0.5× Kimura B (KB) medium for 7 d, followed by 0.4 μmol/L 67Zn re-supply to the leaves via foliar spray or 0.5× KB containing 0.4 μmol/L 67Zn as the local Zn system for 1 h. The control was 0.01% Trition X-100. Values represent Mean ± SD (n ≥ 3). Statistical comparison was performed with the Tukey’s HSD mean-separation test (*, P < 0.05; **, P < 0.01).
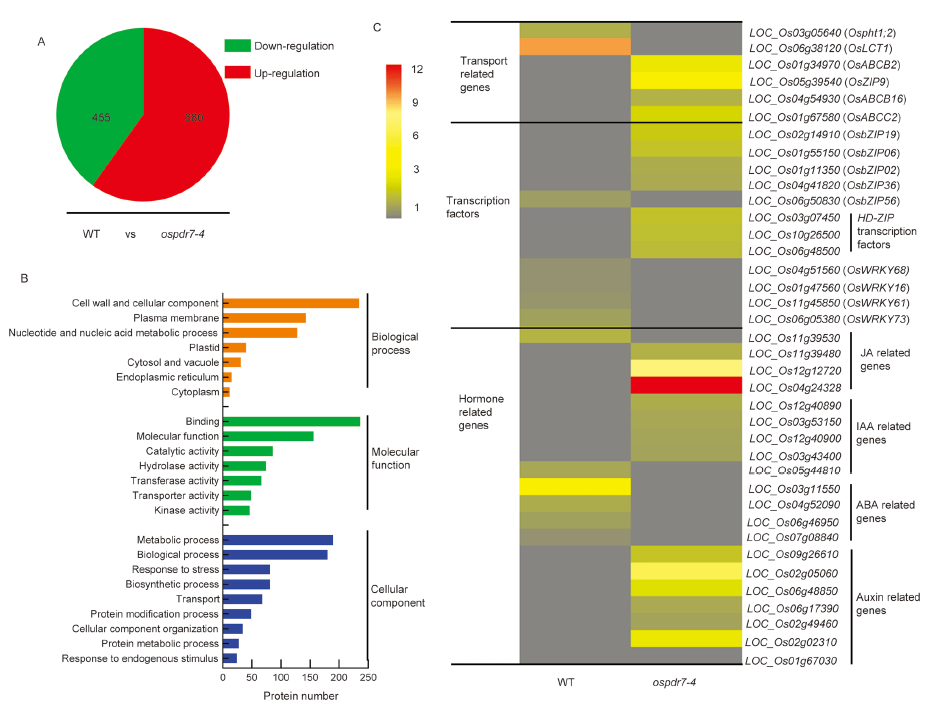
Fig. 5. Differentially expressed genes (DEGs) between wild type (WT) and OsPDR7 mutant (ospdr7-4). A, Number of DEGs between WT and ospdr7-4. B, Gene Ontology enrichment analysis of DEGs (P ≤ 0.05). C, A heat map showing relative expression of transcription factor, transporter, and hormone responsive-related genes. JA, Jasmonic acid; IAA, Indole-3-acetic acid; ABA, Abscisic acid. The roots of WT and ospdr7-4 seedlings (28-day-old) were collected for transcriptome analysis. To identify DEGs, the expression level of each transcript was calculated using the transcripts per million reads method (n = 3).
| [1] |
Assunção A G L, Herrero E, Lin Y F, Huettel B, Talukdar S, Smaczniak C, Immink R G H, van Eldik M, Fiers M, Schat H, Aarts M G M. 2010. Arabidopsis thaliana transcription factors bZIP19 and bZIP23 regulate the adaptation to zinc deficiency. Proc Natl Acad Sci USA, 107(22): 10296-10301.
PMID |
| [2] | Broadley M, Brown P, Cakmak I, Rengel Z, Zhao F J. 2012. Function of nutrients. In: Marschner P. Marschner’s Mineral Nutrition of Higher Plants. Amsterdam, the Netherlands: Elsevier: 191-248. |
| [3] |
Broadley M R, White P J, Hammond J P, Zelko I, Lux A. 2007. Zinc in plants. New Phytol, 173(4): 677-702.
PMID |
| [4] |
Cai H M, Huang S, Che J, Yamaji N, Ma J F. 2019. The tonoplast-localized transporter OsHMA3 plays an important role in maintaining Zn homeostasis in rice. J Exp Bot, 70(10): 2717-2725.
PMID |
| [5] | Cakmak I, Yilmaz A, Kalayci M, Ekiz H, Torun B, Erenoglu B, Braun H J. 1996. Zinc deficiency as a critical nutritional problem in wheat production in Central Anatolia. Plant Soil, 180: 165-172. |
| [6] |
Chen L, Zhao J Q, Song J C, Jameson P E. 2020. Cytokinin dehydrogenase: A genetic target for yield improvement in wheat. Plant Biotechnol J, 18(3): 614-630.
PMID |
| [7] | Chen W R, Feng Y, Chao Y E. 2008. Genomic analysis and expression pattern of OsZIP1, OsZIP3, and OsZIP4 in two rice (Oryza sativa L.) genotypes with different zinc efficiency. Russ J Plant Physiol, 55(3): 400-409. |
| [8] |
Do T H T, Martinoia E, Lee Y. 2018. Functions of ABC transporters in plant growth and development. Curr Opin Plant Biol, 41: 32-38.
PMID |
| [9] | Fang Y, Wang L, Xin Z H, Zhao L Y, An X X, Hu Q H. 2008. Effect of foliar application of zinc, selenium, and iron fertilizers on nutrients concentration and yield of rice grain in China. J Agric Food Chem, 56(6): 2079-2084. |
| [10] |
Fu S, Lu Y S, Zhang X, Yang G Z, Chao D, Wang Z G, Shi M X, Chen J G, Chao D Y, Li R B, Ma J F, Xia J X. 2019. The ABC transporter ABCG36 is required for cadmium tolerance in rice. J Exp Bot, 70(20): 5909-5918.
PMID |
| [11] |
Gao S P, Xiao Y H, Xu F, Gao X K, Cao S Y, Zhang F X, Wang G D, Sanders D, Chu C C. 2019. Cytokinin-dependent regulatory module underlies the maintenance of zinc nutrition in rice. New Phytol, 224(1): 202-215.
PMID |
| [12] | Gupta B B, Selter L L, Baranwal V K, Arora D, Mishra S K, Sirohi P, Poonia A K, Chaudhary R, Kumar R, Krattinger S G, Chauhan H. 2019. Updated inventory, evolutionary and expression analyses of G (PDR) type ABC transporter genes of rice. Plant Physiol Biochem, 142: 429-439. |
| [13] |
Higgins C F, Linton K J. 2004. The ATP switch model for ABC transporters. Nat Struct Mol Biol, 11(10): 918-926.
PMID |
| [14] |
Huang S, Sasaki A, Yamaji N, Okada H, Mitani-Ueno N, Ma J F. 2020. The ZIP transporter family member OsZIP9 contributes to root zinc uptake in rice under zinc-limited conditions. Plant Physiol, 183(3): 1224-1234.
PMID |
| [15] |
Ishimaru Y, Suzuki M, Kobayashi T, Takahashi M, Nakanishi H, Mori S, Nishizawa N K. 2005. OsZIP4, a novel zinc-regulated zinc transporter in rice. J Exp Bot, 56: 3207-3214.
PMID |
| [16] |
Ishimaru Y, Masuda H, Suzuki M, Bashir K, Takahashi M, Nakanishi H, Mori S, Nishizawa N K. 2007. Overexpression of the OsZIP4 zinc transporter confers disarrangement of zinc distribution in rice plants. J Exp Bot, 58(11): 2909-2915.
PMID |
| [17] | Ito H, Gray W M. 2006. A gain-of-function mutation in the Arabidopsis pleiotropic drug resistance transporter PDR9 confers resistance to auxinic herbicides. Plant Physiol, 142(1): 63-74. |
| [18] |
Jakoby M, Weisshaar B, Dröge-Laser W, Vicente-Carbajosa J, Tiedemann J, Kroj T, Parcy F, Group B R. 2002. bZIP transcription factors in Arabidopsis. Trends Plant Sci, 7(3): 106-111.
PMID |
| [19] |
Kang J, Hwang J U, Lee M, Kim Y Y, Assmann S M, Martinoia E, Lee Y. 2010. PDR-type ABC transporter mediates cellular uptake of the phytohormone abscisic acid. Proc Natl Acad Sci USA, 107(5): 2355-2360.
PMID |
| [20] | Kawakami Y, Bhullar N K. 2018. Molecular processes in iron and zinc homeostasis and their modulation for biofortification in rice. J Integr Plant Biol, 60(12): 1181-1198. |
| [21] | Kim D Y, Bovet L, Maeshima M, Martinoia E, Lee Y. 2007. The ABC transporter AtPDR8 is a cadmium extrusion pump conferring heavy metal resistance. Plant J, 50(2): 207-218. |
| [22] | Kim D Y, Jin J Y, Alejandro S, Martinoia E, Lee Y. 2010. Overexpression of AtABCG 36 improves drought and salt stress resistance in Arabidopsis. Physiol Plant, 139(2): 170-180. |
| [23] | Ko D, Kang J, Kiba T, Park J, Kojima M, Do J, Kim K Y, Kwon M, Endler A, Song W Y, Martinoia E, Sakakibara H, Lee Y. 2014. Arabidopsis ABCG14 is essential for the root-to-shoot translocation of cytokinin. Proc Natl Acad Sci USA, 111(19): 7150-7155. |
| [24] |
Kuromori T, Miyaji T, Yabuuchi H, Shimizu H, Sugimoto E, Kamiya A, Moriyama Y, Shinozaki K. 2010. ABC transporter AtABCG25 is involved in abscisic acid transport and responses. Proc Natl Acad Sci USA, 107(5): 2361-2366.
PMID |
| [25] |
Lee M, Lee K, Lee J, Noh E W, Lee Y. 2005. AtPDR12 contributes to lead resistance in Arabidopsis. Plant Physiol, 138(2): 827-836.
PMID |
| [26] | Lei G J, Sun L, Sun Y, Zhu X F, Li G X, Zheng S J. 2020. Jasmonic acid alleviates cadmium toxicity in Arabidopsis via suppression of cadmium uptake and translocation. J Integr Plant Biol, 62(2): 218-227. |
| [27] |
Lilay G H, Castro P H, Guedes J G, Almeida D M, Campilho A, Azevedo H, Aarts M G M, Saibo N J M, Assunção A G L. 2020. Rice F-bZIP transcription factors regulate the zinc deficiency response. J Exp Bot, 71(12): 3664-3677.
PMID |
| [28] | Love M I, Huber W, Anders S. 2014. Moderated estimation of fold change and dispersion for RNA-Seq data with DESeq2. Genome Biol, 15(12): 550. |
| [29] |
Lu Y M, Ye X, Guo R M, Huang J, Wang W, Tang J Y, Tan L T, Zhu J K, Chu C C, Qian Y W. 2017. Genome-wide targeted mutagenesis in rice using the CRISPR/Cas9 system. Mol Plant, 10(9): 1242-1245.
PMID |
| [30] |
Menguer P K, Farthing E, Peaston K A, Ricachenevsky F K, Fett J P, Williams L E. 2013. Functional analysis of the rice vacuolar zinc transporter OsMTP1. J Exp Bot, 64(10): 2871-2883.
PMID |
| [31] |
Miyao A, Tanaka K, Murata K, Sawaki H, Takeda S, Abe K, Shinozuka Y, Onosato K, Hirochika H. 2003. Target site specificity of the Tos17 retrotransposon shows a preference for insertion within genes and against insertion in retrotransposon- rich regions of the genome. Plant Cell, 15(8): 1771-1780.
PMID |
| [32] | Mu S, Yamaji N, Sasaki A, Luo L, Du B B, Che J, Shi H C, Zhao H Q, Huang S, Deng F L, Shen Z G, Guerinot M L, Zheng L Q, Ma J F. 2021. A transporter for delivering zinc to the developing tiller bud and panicle in rice. Plant J, 105(3): 786-799. |
| [33] | Muvunyi B P, Lu X, Zhan J H, He S, Ye G Y. 2022. Identification of potential zinc deficiency responsive genes and regulatory pathways in rice by weighted gene co-expression network analysis. Rice Sci, 29(6): 545-558. |
| [34] | Oda K, Otani M, Uraguchi S, Akihiro T, Fujiwara T. 2011. Rice ABCG43 is Cd inducible and confers Cd tolerance on yeast. Biosci Biotechnol Biochem, 75(6): 1211-1213. |
| [35] | Park J, Song W Y, Ko D, Eom Y, Hansen T H, Schiller M, Lee T G, Martinoia E, Lee Y. 2012. The phytochelatin transporters AtABCC1 and AtABCC2 mediate tolerance to cadmium and mercury. Plant J, 69(2): 278-288. |
| [36] |
Patro R, Duggal G, Love M I, Irizarry R A, Kingsford C. 2017. Salmon provides fast and bias-aware quantification of transcript expression. Nat Methods, 14(4): 417-419.
PMID |
| [37] |
Rea P A. 2007. Plant ATP-binding cassette transporters. Annu Rev Plant Biol, 58: 347-375.
PMID |
| [38] | Sasaki A, Yamaji N, Mitani-Ueno N, Kashino M, Ma J F. 2015. A node-localized transporter OsZIP3 is responsible for the preferential distribution of Zn to developing tissues in rice. Plant J, 84(2): 374-384. |
| [39] | Song C Z, Yan Y F, Rosado A, Zhang Z W, Castellarin S D. 2019. ABA alleviates uptake and accumulation of zinc in grapevine (Vitis vinifera L.) by inducing expression of ZIP and detoxification- related genes. Front Plant Sci, 10: 872. |
| [40] |
Sparkes I A, Runions J, Kearns A, Hawes C. 2006. Rapid, transient expression of fluorescent fusion proteins in tobacco plants and generation of stably transformed plants. Nat Protoc, 1(4): 2019-2025.
PMID |
| [41] |
Stein M, Dittgen J, Sánchez-Rodríguez C, Hou B H, Molina A, Schulze-Lefert P, Lipka V, Somerville S. 2006. Arabidopsis PEN3/PDR8, an ATP binding cassette transporter, contributes to nonhost resistance to inappropriate pathogens that enter by direct penetration. Plant Cell, 18(3): 731-746.
PMID |
| [42] |
Swamy B P M, Rahman M A, Inabangan-Asilo M A, Amparado A, Manito C, Chadha-Mohanty P, Reinke R, Slamet-Loedin I H. 2016. Advances in breeding for high grain zinc in rice. Rice, 9(1): 49.
PMID |
| [43] | Takahashi R, Ishimaru Y, Shimo H, Ogo Y, Senoura T, Nishizawa N K, Nakanishi H. 2012. The OsHMA2 transporter is involved in root-to-shoot translocation of Zn and Cd in rice. Plant Cell Environ, 35(11): 1948-1957. |
| [44] | Tan L T, Zhu Y X, Fan T, Peng C, Wang J R, Sun L, Chen C Y. 2019. OsZIP7 functions in xylem loading in roots and inter- vascular transfer in nodes to deliver Zn/Cd to grain in rice. Biochem Biophys Res Commun, 512(1): 112-118. |
| [45] |
Tan L T, Qu M M, Zhu Y X, Peng C, Wang J R, Gao D Y, Chen C Y. 2020. ZINC TRANSPORTER5 and ZINC TRANSPORTER9 function synergistically in zinc/cadmium uptake. Plant Physiol, 183(3): 1235-1249.
PMID |
| [46] | Tong J Y, Sun M J, Wang Y, Zhang Y, Rasheed A, Li M, Xia X C, He Z H, Hao Y F. 2020. Dissection of molecular processes and genetic architecture underlying iron and zinc homeostasis for biofortification: From model plants to common wheat. Int J Mol Sci, 21(23): 9280. |
| [47] | Utasee S, Jamjod S, Lordkaew S, Prom-U-Thai C. 2022. Improve anthocyanin and zinc concentration in purple rice by nitrogen and zinc fertilizer application. Rice Sci, 29(5): 435-450. |
| [48] |
Verrier P J, Bird D, Burla B, Dassa E, Forestier C, Geisler M, Klein M, Kolukisaoglu U, Lee Y, Martinoia E, Murphy A, Rea P A, Samuels L, Schulz B, Spalding E J, Yazaki K, Theodoulou F L. 2008. Plant ABC proteins: A unified nomenclature and updated inventory. Trends Plant Sci, 13(4): 151-159.
PMID |
| [49] | Wang Z Y, Wang H, Xu C, Lv G H, Luo Z C, Zhu H H, Wang S, Zhu Q H, Huang D Y, Li B Z. 2020. Foliar application of Zn-EDTA at early filling stage to increase grain Zn and Fe, and reduce grain Cd, Pb and grain yield in rice (Oryza sativa L.). Bull Environ Contam Toxicol, 105(3): 428-432. |
| [50] |
Yamaji N, Ma J F. 2014. The node, a hub for mineral nutrient distribution in graminaceous plants. Trends Plant Sci, 19(9): 556-563.
PMID |
| [51] |
Yamaji N, Xia J X, Mitani-Ueno N, Yokosho K, Ma J F. 2013. Preferential delivery of zinc to developing tissues in rice is mediated by P-type heavy metal ATPase OsHMA2. Plant Physiol, 162(2): 927-939.
PMID |
| [52] | Yang M, Li Y T, Liu Z H, Tian J J, Liang L M, Qiu Y, Wang G Y, Du Q Q, Cheng D, Cai H M, Shi L, Xu F S, Lian X M. 2020. A high activity zinc transporter OsZIP9 mediates zinc uptake in rice. Plant J, 103(5): 1695-1709. |
| [53] | Yoo S D, Cho Y H, Sheen J. 2007. Arabidopsis mesophyll protoplasts: A versatile cell system for transient gene expression analysis. Nat Protoc, 2(7): 1565-1572. |
| [54] | Zhang H S, Jing W, Zheng J M, Jin Y Y, Wu D, Cao C J, Dong Y M, Shi X Y, Zhang W H. 2020. The ATP-binding cassette transporter OsPDR1 regulates plant growth and pathogen resistance by affecting jasmonates biosynthesis in rice. Plant Sci, 298: 110582. |
| [55] | Zhang K W, Novak O, Wei Z Y, Gou M Y, Zhang X B, Yu Y, Yang H J, Cai Y H, Strnad M, Liu C J. 2014. Arabidopsis ABCG14 protein controls the acropetal translocation of root-synthesized cytokinins. Nat Commun, 5: 3274. |
| [56] |
Zhao J Z, Yu N N, Ju M, Fan B, Zhang Y J, Zhu E G, Zhang M Y, Zhang K W. 2019. ABC transporter OsABCG18 controls the shootward transport of cytokinins and grain yield in rice. J Exp Bot, 70(21): 6277-6291.
PMID |
| [1] | Prathap V, Suresh KUMAR, Nand Lal MEENA, Chirag MAHESHWARI, Monika DALAL, Aruna TYAGI. Phosphorus Starvation Tolerance in Rice Through a Combined Physiological, Biochemical and Proteome Analysis [J]. Rice Science, 2023, 30(6): 8-. |
| [2] | Serena REGGI, Elisabetta ONELLI, Alessandra MOSCATELLI, Nadia STROPPA, Matteo Dell’ANNO, Kiril PERFANOV, Luciana ROSSI. Seed-Specific Expression of Apolipoprotein A-IMilano Dimer in Rice Engineered Lines [J]. Rice Science, 2023, 30(6): 6-. |
| [3] | Sundus ZAFAR, XU Jianlong. Recent Advances to Enhance Nutritional Quality of Rice [J]. Rice Science, 2023, 30(6): 4-. |
| [4] | Kankunlanach KHAMPUANG, Nanthana CHAIWONG, Atilla YAZICI, Baris DEMIRER, Ismail CAKMAK, Chanakan PROM-U-THAI. Effect of Sulfur Fertilization on Productivity and Grain Zinc Yield of Rice Grown under Low and Adequate Soil Zinc Applications [J]. Rice Science, 2023, 30(6): 9-. |
| [5] | FAN Fengfeng, CAI Meng, LUO Xiong, LIU Manman, YUAN Huanran, CHENG Mingxing, Ayaz AHMAD, LI Nengwu, LI Shaoqing. Novel QTLs from Wild Rice Oryza longistaminata Confer Rice Strong Tolerance to High Temperature at Seedling Stage [J]. Rice Science, 2023, 30(6): 14-. |
| [6] | LIN Shaodan, YAO Yue, LI Jiayi, LI Xiaobin, MA Jie, WENG Haiyong, CHENG Zuxin, YE Dapeng. Application of UAV-Based Imaging and Deep Learning in Assessment of Rice Blast Resistance [J]. Rice Science, 2023, 30(6): 10-. |
| [7] | Md. Forshed DEWAN, Md. AHIDUZZAMAN, Md. Nahidul ISLAM, Habibul Bari SHOZIB. Potential Benefits of Bioactive Compounds of Traditional Rice Grown in South and South-East Asia: A Review [J]. Rice Science, 2023, 30(6): 5-. |
| [8] | Raja CHAKRABORTY, Pratap KALITA, Saikat SEN. Phenolic Profile, Antioxidant, Antihyperlipidemic and Cardiac Risk Preventive Effect of Chakhao Poireiton (A Pigmented Black Rice) in High-Fat High-Sugar induced Rats [J]. Rice Science, 2023, 30(6): 11-. |
| [9] | LI Qianlong, FENG Qi, WANG Heqin, KANG Yunhai, ZHANG Conghe, DU Ming, ZHANG Yunhu, WANG Hui, CHEN Jinjie, HAN Bin, FANG Yu, WANG Ahong. Genome-Wide Dissection of Quan 9311A Breeding Process and Application Advantages [J]. Rice Science, 2023, 30(6): 7-. |
| [10] | JI Dongling, XIAO Wenhui, SUN Zhiwei, LIU Lijun, GU Junfei, ZHANG Hao, Tom Matthew HARRISON, LIU Ke, WANG Zhiqin, WANG Weilu, YANG Jianchang. Translocation and Distribution of Carbon-Nitrogen in Relation to Rice Yield and Grain Quality as Affected by High Temperature at Early Panicle Initiation Stage [J]. Rice Science, 2023, 30(6): 12-. |
| [11] | Nazaratul Ashifa Abdullah Salim, Norlida Mat Daud, Julieta Griboff, Abdul Rahim Harun. Elemental Assessments in Paddy Soil for Geographical Traceability of Rice from Peninsular Malaysia [J]. Rice Science, 2023, 30(5): 486-498. |
| [12] | Monica Ruffini Castiglione, Stefania Bottega, Carlo Sorce, Carmelina SpanÒ. Effects of Zinc Oxide Particles with Different Sizes on Root Development in Oryza sativa [J]. Rice Science, 2023, 30(5): 449-458. |
| [13] | Tan Jingyi, Zhang Xiaobo, Shang Huihui, Li Panpan, Wang Zhonghao, Liao Xinwei, Xu Xia, Yang Shihua, Gong Junyi, Wu Jianli. ORYZA SATIVA SPOTTED-LEAF 41 (OsSPL41) Negatively Regulates Plant Immunity in Rice [J]. Rice Science, 2023, 30(5): 426-436. |
| [14] | Ammara Latif, Sun Ying, Pu Cuixia, Noman Ali. Rice Curled Its Leaves Either Adaxially or Abaxially to Combat Drought Stress [J]. Rice Science, 2023, 30(5): 405-416. |
| [15] | Liu Qiao, Qiu Linlin, Hua Yangguang, Li Jing, Pang Bo, Zhai Yufeng, Wang Dekai. LHD3 Encoding a J-Domain Protein Controls Heading Date in Rice [J]. Rice Science, 2023, 30(5): 437-448. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||