
Rice Science ›› 2020, Vol. 27 ›› Issue (6): 449-453.DOI: 10.1016/j.rsci.2020.09.002
• Letter • Previous Articles Next Articles
Kaur Sandhu Rajwinder1, Singh Sarao Preetinder2( ), Kumari Neelam3
), Kumari Neelam3
Received:2019-09-13
Accepted:2020-03-25
Online:2020-11-28
Published:2020-11-28
Kaur Sandhu Rajwinder, Singh Sarao Preetinder, Kumari Neelam. Biochemical Responses Associated with Resistance to Nilaparvata lugens (Stål) in Wild Rice Accessions[J]. Rice Science, 2020, 27(6): 449-453.
Add to citation manager EndNote|Ris|BibTeX
| Accession | Modified seed box score | Categorization on the basis of SES | Excreted honeydew area (mm2) | Weight of filter paper with honeydew (mg) | Watery lesion |
|---|---|---|---|---|---|
| IRGC99577 (O. punctata) | 2.17 ± 0.30 | Resistant | 74.10 ± 1.10 a | 9.50 ± 0.30 a | 2.70 ± 0.10 e |
| IRGC105270 (O. australiensis) | 3.43 ± 0.43 | Resistant | 101.50 ± 1.10 b | 13.50 ± 0.50 b | 2.50 ± 0.10 de |
| IRGC105275 (O. australiensis) | 4.60 ± 0.40 | Moderately resistant | 123.70 ± 1.30 c | 16.40 ± 0.20 c | 2.10 ± 0.10 c |
| IRGC104646 (O. nivara) | 2.85 ± 0.40 | Resistant | 113.90 ± 1.90 c | 14.80 ± 0.40 bc | 2.30 ± 0.10 cd |
| CR100204 (O. nivara) | 5.03 ± 0.30 | Moderately resistant | 147.90 ± 6.30 d | 20.00 ± 1.20 d | 1.70 ± 0.10 b |
| Ptb33 (Resistant check) | 2.30 ± 0.30 | Resistant | 78.60 ± 1.20 a | 10.10 ± 0.30 a | 2.60 ± 0.00 de |
| TN1 (Susceptible check) | 9.00 ± 0.00 | Susceptible | 412.70 ± 4.30 e | 52.40 ± 1.00 e | 0.30 ± 0.10 a |
| LSD0.05 | 1.12 | 0.84 | 2.21 | 0.31 | |
| SES, Standard evaluation system. Data are Mean ± SE (n = 30 for modified seed box score and 5 for honeydew area and watery lesion). Means within a column followed by the same lowercase letters are not significantly different at P ≤ 0.05 according to the Duncan’s multiple range test. | |||||
Table 1 Response of wild rice accessions to Nilaparvata lugens.
| Accession | Modified seed box score | Categorization on the basis of SES | Excreted honeydew area (mm2) | Weight of filter paper with honeydew (mg) | Watery lesion |
|---|---|---|---|---|---|
| IRGC99577 (O. punctata) | 2.17 ± 0.30 | Resistant | 74.10 ± 1.10 a | 9.50 ± 0.30 a | 2.70 ± 0.10 e |
| IRGC105270 (O. australiensis) | 3.43 ± 0.43 | Resistant | 101.50 ± 1.10 b | 13.50 ± 0.50 b | 2.50 ± 0.10 de |
| IRGC105275 (O. australiensis) | 4.60 ± 0.40 | Moderately resistant | 123.70 ± 1.30 c | 16.40 ± 0.20 c | 2.10 ± 0.10 c |
| IRGC104646 (O. nivara) | 2.85 ± 0.40 | Resistant | 113.90 ± 1.90 c | 14.80 ± 0.40 bc | 2.30 ± 0.10 cd |
| CR100204 (O. nivara) | 5.03 ± 0.30 | Moderately resistant | 147.90 ± 6.30 d | 20.00 ± 1.20 d | 1.70 ± 0.10 b |
| Ptb33 (Resistant check) | 2.30 ± 0.30 | Resistant | 78.60 ± 1.20 a | 10.10 ± 0.30 a | 2.60 ± 0.00 de |
| TN1 (Susceptible check) | 9.00 ± 0.00 | Susceptible | 412.70 ± 4.30 e | 52.40 ± 1.00 e | 0.30 ± 0.10 a |
| LSD0.05 | 1.12 | 0.84 | 2.21 | 0.31 | |
| SES, Standard evaluation system. Data are Mean ± SE (n = 30 for modified seed box score and 5 for honeydew area and watery lesion). Means within a column followed by the same lowercase letters are not significantly different at P ≤ 0.05 according to the Duncan’s multiple range test. | |||||
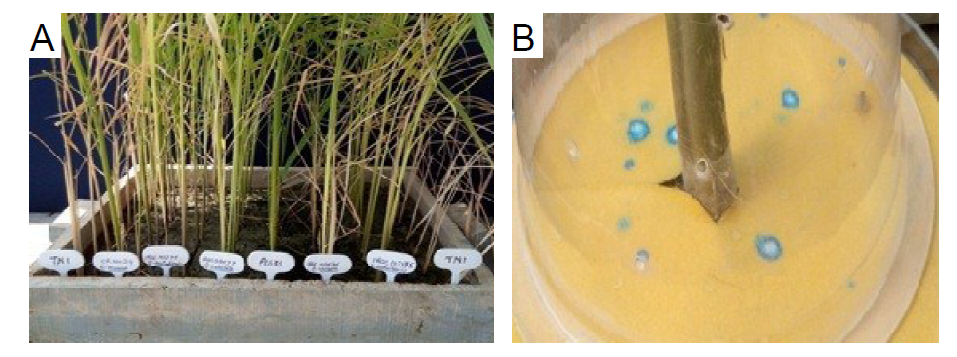

Supplemental Fig. 1. Antibiosis resistance parameters. A, Modified seed box screening (MSBS) of wild rice accessions. B, Honeydew excretion by five adult females of Nilaparvata lugens on filter paper.
| Accession | PAL [μg/(h·mg)] | PPO [U/(min·mg)] | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Leaf blade | Leaf sheath | Leaf blade | Leaf sheath | ||||||||
| Constitutive level | Induced level | Constitutive level | Induced level | Constitutive level | Induced level | Constitutive level | Induced level | ||||
| IRGC99577 | 2.23 ± 0.05 e | 2.36 ± 0.05 c | 1.91 ± 0.04 d | 2.16 ± 0.07 d | 0.19 ± 0.00 e | 0.21 ± 0.01 e | 0.16 ± 0.01 d | 0.18 ± 0.01 d | |||
| IRGC105270 | 2.02 ± 0.05 d | 2.17 ± 0.05 b | 1.80 ± 0.03 c | 1.94 ± 0.06 c | 0.15 ± 0.02 cde | 0.17 ± 0.01 cd | 0.11 ± 0.00 c | 0.13 ± 0.01 bc | |||
| IRGC105275 | 1.80 ± 0.03 bc | 1.92 ± 0.02 a | 1.51 ± 0.02 b | 1.65 ± 0.04 b | 0.13 ± 0.01 bc | 0.15 ± 0.00 c | 0.08 ± 0.00 b | 0.12 ± 0.01 bc | |||
| IRGC104646 | 1.93 ± 0.06 cd | 2.11 ± 0.06 b | 1.71 ± 0.04 c | 1.84 ± 0.03 c | 0.15 ± 0.01 cd | 0.17 ± 0.01 c | 0.12 ± 0.01 c | 0.13 ± 0.01 c | |||
| CR100204 | 1.75 ± 0.04 b | 1.87 ± 0.04 a | 1.45 ± 0.03 b | 1.60 ± 0.04 b | 0.11 ± 0.01 b | 0.13 ± 0.01 b | 0.08 ± 0.00 b | 0.09 ± 0.01 ab | |||
| Ptb33 | 2.25 ± 0.04 e | 2.40 ± 0.04 c | 1.93 ± 0.03 d | 2.20 ± 0.05 d | 0.17 ± 0.01 de | 0.19 ± 0.01 de | 0.14 ± 0.01 d | 0.17 ± 0.01 d | |||
| TN1 | 1.57 ± 0.02 a | 1.76 ± 0.04 a | 1.30 ± 0.02 a | 1.39 ± 0.01 a | 0.07 ± 0.01 a | 0.09 ± 0.00 a | 0.05 ± 0.00 a | 0.06 ± 0.01 a | |||
| LSD0.05 | |||||||||||
| Accession (A) | 0.05 | 0.04 | 0.09 | 0.09 | |||||||
| Infestation (I) | 0.10 | 0.08 | 0.02 | 0.18 | |||||||
| A × I | Not significant | Not significant | Not significant | Not significant | |||||||
| Values are Mean ± SE (n = 3). Means within a column followed by the same lowercase letters are not significantly different at P ≤ 0.05 according to the Duncan’s multiple range test. | |||||||||||
Table 2 Specific activities of phenylalanine ammonia lyase (PAL) and polyphenol oxidase (PPO) in leaf blade and leaf sheath of rice accessions at constitutive and induced levels against N. lugens.
| Accession | PAL [μg/(h·mg)] | PPO [U/(min·mg)] | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Leaf blade | Leaf sheath | Leaf blade | Leaf sheath | ||||||||
| Constitutive level | Induced level | Constitutive level | Induced level | Constitutive level | Induced level | Constitutive level | Induced level | ||||
| IRGC99577 | 2.23 ± 0.05 e | 2.36 ± 0.05 c | 1.91 ± 0.04 d | 2.16 ± 0.07 d | 0.19 ± 0.00 e | 0.21 ± 0.01 e | 0.16 ± 0.01 d | 0.18 ± 0.01 d | |||
| IRGC105270 | 2.02 ± 0.05 d | 2.17 ± 0.05 b | 1.80 ± 0.03 c | 1.94 ± 0.06 c | 0.15 ± 0.02 cde | 0.17 ± 0.01 cd | 0.11 ± 0.00 c | 0.13 ± 0.01 bc | |||
| IRGC105275 | 1.80 ± 0.03 bc | 1.92 ± 0.02 a | 1.51 ± 0.02 b | 1.65 ± 0.04 b | 0.13 ± 0.01 bc | 0.15 ± 0.00 c | 0.08 ± 0.00 b | 0.12 ± 0.01 bc | |||
| IRGC104646 | 1.93 ± 0.06 cd | 2.11 ± 0.06 b | 1.71 ± 0.04 c | 1.84 ± 0.03 c | 0.15 ± 0.01 cd | 0.17 ± 0.01 c | 0.12 ± 0.01 c | 0.13 ± 0.01 c | |||
| CR100204 | 1.75 ± 0.04 b | 1.87 ± 0.04 a | 1.45 ± 0.03 b | 1.60 ± 0.04 b | 0.11 ± 0.01 b | 0.13 ± 0.01 b | 0.08 ± 0.00 b | 0.09 ± 0.01 ab | |||
| Ptb33 | 2.25 ± 0.04 e | 2.40 ± 0.04 c | 1.93 ± 0.03 d | 2.20 ± 0.05 d | 0.17 ± 0.01 de | 0.19 ± 0.01 de | 0.14 ± 0.01 d | 0.17 ± 0.01 d | |||
| TN1 | 1.57 ± 0.02 a | 1.76 ± 0.04 a | 1.30 ± 0.02 a | 1.39 ± 0.01 a | 0.07 ± 0.01 a | 0.09 ± 0.00 a | 0.05 ± 0.00 a | 0.06 ± 0.01 a | |||
| LSD0.05 | |||||||||||
| Accession (A) | 0.05 | 0.04 | 0.09 | 0.09 | |||||||
| Infestation (I) | 0.10 | 0.08 | 0.02 | 0.18 | |||||||
| A × I | Not significant | Not significant | Not significant | Not significant | |||||||
| Values are Mean ± SE (n = 3). Means within a column followed by the same lowercase letters are not significantly different at P ≤ 0.05 according to the Duncan’s multiple range test. | |||||||||||
| Accession | Total phenol (mg/g) | Flavonol (mg/g) | Total free amino acid (mg/g) | ||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Leaf blade | Leaf sheath | Leaf blade | Leaf sheath | Leaf blade | Leaf sheath | ||||||||||||
| Constitutive level | Induced level | Constitutive level | Induced level | Constitutive level | Induced level | Constitutive level | Induced level | Constitutive level | Induced level | Constitutive level | Induced level | ||||||
| IRGC99577 | 6.18 ± 0.06 d | 6.36 ± 0.05 d | 2.63 ± 0.08 d | 2.87 ± 0.03 c | 2.49 ± 0.07 e | 2.97 ± 0.06 e | 0.73 ± 0.07 ab | 1.27 ± 0.04 b | 1.15 ± 0.12 a | 1.30 ± 0.12 a | 1.11 ± 0.10 a | 1.26 ± 0.13 a | |||||
| IRGC105270 | 5.58 ± 0.09 c | 5.94 ± 0.05 c | 2.16 ± 0.06 c | 2.72 ± 0.04 c | 1.85 ± 0.13 cd | 2.61 ± 0.07 d | 0.67 ± 0.08 ab | 1.08 ± 0.16 ab | 1.44 ± 0.12 abc | 1.69 ± 0.11 abc | 1.23 ± 0.11 a | 1.44 ± 0.15 ab | |||||
| IRGC105275 | 5.06 ± 0.06 b | 5.30 ± 0.04 b | 1.87 ± 0.04 b | 2.25 ± 0.07 b | 1.68 ± 0.07 bc | 1.99 ± 0.06 b | 0.62 ± 0.07 ab | 0.95 ± 0.09 ab | 1.53 ± 0.09 bc | 1.80 ± 0.11 bcd | 1.25 ± 0.11 a | 1.47 ± 0.16 ab | |||||
| IRGC104646 | 5.97 ± 0.04 d | 6.21 ± 0.06 d | 2.18 ± 0.06 c | 2.78 ± 0.03 c | 2.02 ± 0.04 d | 2.34 ± 0.05 c | 0.64 ± 0.07 ab | 1.02 ± 0.13 ab | 1.44 ± 0.11 abc | 1.67 ± 0.13 abc | 1.22 ± 0.10 a | 1.42 ± 0.13 ab | |||||
| CR100204 | 4.61 ± 0.07 a | 5.16 ± 0.07 b | 1.79 ± 0.07 b | 2.09 ± 0.07 b | 1.49 ± 0.14 b | 1.87 ± 0.13 b | 0.57 ± 0.05 ab | 0.82 ± 0.08 a | 1.52 ± 0.07 bc | 1.85 ± 0.13 cd | 1.31 ± 0.08 a | 1.56 ± 0.12 ab | |||||
| Ptb33 | 7.62 ± 0.11 e | 7.79 ± 0.13 e | 2.72 ± 0.07 d | 3.77 ± 0.05 d | 2.75 ± 0.06 e | 3.07 ± 0.06 e | 0.78 ± 0.07 b | 1.30 ± 0.06 b | 1.24 ± 0.07 ab | 1.40 ± 0.08 ab | 1.16 ± 0.06 a | 1.32 ± 0.10 ab | |||||
| TN1 | 4.35 ± 0.10 a | 4.90 ± 0.03 a | 1.42 ± 0.09 a | 1.69 ± 0.06 a | 1.15 ± 0.08 a | 1.52 ± 0.07 a | 0.52 ± 0.05 a | 0.83 ± 0.11 a | 1.70 ± 0.09 c | 2.14 ± 0.13 d | 1.42 ± 0.11 a | 1.76 ± 0.13 b | |||||
| LSD (P = 0.05) | |||||||||||||||||
| Accession | 0.08 | 0.07 | 0.10 | 0.10 | 0.12 | 0.13 | |||||||||||
| Infestation | 0.16 | 0.13 | 0.18 | 0.19 | 0.22 | NS | |||||||||||
| Accession × Infestation | NS | 0.19 | NS | NS | NS | NS | |||||||||||
Supplemental Table 1. Total phenol, flavonol and total free amino acid contents in leaf blade and leaf sheath of rice accessions at constitutive and induced levels against N. lugens.
| Accession | Total phenol (mg/g) | Flavonol (mg/g) | Total free amino acid (mg/g) | ||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Leaf blade | Leaf sheath | Leaf blade | Leaf sheath | Leaf blade | Leaf sheath | ||||||||||||
| Constitutive level | Induced level | Constitutive level | Induced level | Constitutive level | Induced level | Constitutive level | Induced level | Constitutive level | Induced level | Constitutive level | Induced level | ||||||
| IRGC99577 | 6.18 ± 0.06 d | 6.36 ± 0.05 d | 2.63 ± 0.08 d | 2.87 ± 0.03 c | 2.49 ± 0.07 e | 2.97 ± 0.06 e | 0.73 ± 0.07 ab | 1.27 ± 0.04 b | 1.15 ± 0.12 a | 1.30 ± 0.12 a | 1.11 ± 0.10 a | 1.26 ± 0.13 a | |||||
| IRGC105270 | 5.58 ± 0.09 c | 5.94 ± 0.05 c | 2.16 ± 0.06 c | 2.72 ± 0.04 c | 1.85 ± 0.13 cd | 2.61 ± 0.07 d | 0.67 ± 0.08 ab | 1.08 ± 0.16 ab | 1.44 ± 0.12 abc | 1.69 ± 0.11 abc | 1.23 ± 0.11 a | 1.44 ± 0.15 ab | |||||
| IRGC105275 | 5.06 ± 0.06 b | 5.30 ± 0.04 b | 1.87 ± 0.04 b | 2.25 ± 0.07 b | 1.68 ± 0.07 bc | 1.99 ± 0.06 b | 0.62 ± 0.07 ab | 0.95 ± 0.09 ab | 1.53 ± 0.09 bc | 1.80 ± 0.11 bcd | 1.25 ± 0.11 a | 1.47 ± 0.16 ab | |||||
| IRGC104646 | 5.97 ± 0.04 d | 6.21 ± 0.06 d | 2.18 ± 0.06 c | 2.78 ± 0.03 c | 2.02 ± 0.04 d | 2.34 ± 0.05 c | 0.64 ± 0.07 ab | 1.02 ± 0.13 ab | 1.44 ± 0.11 abc | 1.67 ± 0.13 abc | 1.22 ± 0.10 a | 1.42 ± 0.13 ab | |||||
| CR100204 | 4.61 ± 0.07 a | 5.16 ± 0.07 b | 1.79 ± 0.07 b | 2.09 ± 0.07 b | 1.49 ± 0.14 b | 1.87 ± 0.13 b | 0.57 ± 0.05 ab | 0.82 ± 0.08 a | 1.52 ± 0.07 bc | 1.85 ± 0.13 cd | 1.31 ± 0.08 a | 1.56 ± 0.12 ab | |||||
| Ptb33 | 7.62 ± 0.11 e | 7.79 ± 0.13 e | 2.72 ± 0.07 d | 3.77 ± 0.05 d | 2.75 ± 0.06 e | 3.07 ± 0.06 e | 0.78 ± 0.07 b | 1.30 ± 0.06 b | 1.24 ± 0.07 ab | 1.40 ± 0.08 ab | 1.16 ± 0.06 a | 1.32 ± 0.10 ab | |||||
| TN1 | 4.35 ± 0.10 a | 4.90 ± 0.03 a | 1.42 ± 0.09 a | 1.69 ± 0.06 a | 1.15 ± 0.08 a | 1.52 ± 0.07 a | 0.52 ± 0.05 a | 0.83 ± 0.11 a | 1.70 ± 0.09 c | 2.14 ± 0.13 d | 1.42 ± 0.11 a | 1.76 ± 0.13 b | |||||
| LSD (P = 0.05) | |||||||||||||||||
| Accession | 0.08 | 0.07 | 0.10 | 0.10 | 0.12 | 0.13 | |||||||||||
| Infestation | 0.16 | 0.13 | 0.18 | 0.19 | 0.22 | NS | |||||||||||
| Accession × Infestation | NS | 0.19 | NS | NS | NS | NS | |||||||||||
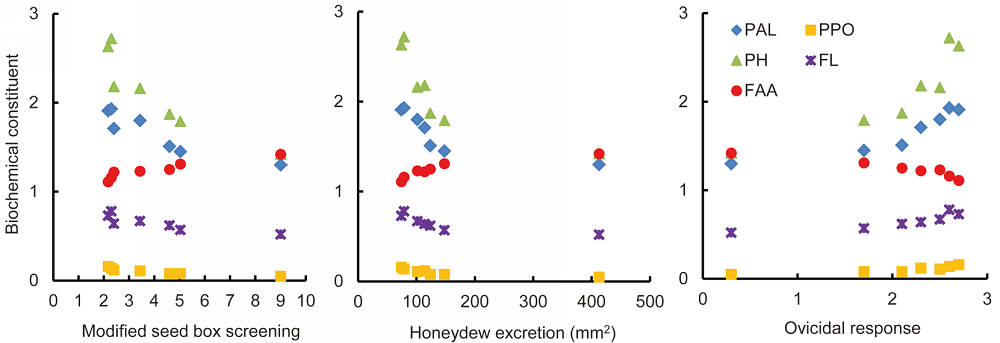
Fig. 1. Pearson’s correlation between biochemical constituents and antibiosis resistance parameters.PAL, Phenylalanine ammonia lyase; PPO, Polyphenol oxidase; PH, Total phenols; FL, Flavonols; FAA, Free amino acids.
| Regression equation | Coefficient of determination (R2) |
|---|---|
| Y1 = -51.05 - 17.39X1 + 208.48X2 - 24.87X3 + 105.65X4 + 37.08X5 | 0.99 |
| Y2 = -2156.32 + 193.31X3 - 267.66X4 + 1667.12X5 | 0.81 |
| Y3 = 19.06 - 18.75X2 + 1.14X3 - 3.29X4 - 12.32X5 | 0.92 |
| Y1 = Damage score, Y2 = Honeydew excretion, Y3 = Ovicidal response, X1 = Phenylalanine ammonia lyase, X2 = Polyphenol oxidase, X3 = Total phenols, X4 = Flavonols, X5 = Free amino acids. | |
Supplemental Table 2. Linear regression analysis between antibiosis parameters and biochemical constituents.
| Regression equation | Coefficient of determination (R2) |
|---|---|
| Y1 = -51.05 - 17.39X1 + 208.48X2 - 24.87X3 + 105.65X4 + 37.08X5 | 0.99 |
| Y2 = -2156.32 + 193.31X3 - 267.66X4 + 1667.12X5 | 0.81 |
| Y3 = 19.06 - 18.75X2 + 1.14X3 - 3.29X4 - 12.32X5 | 0.92 |
| Y1 = Damage score, Y2 = Honeydew excretion, Y3 = Ovicidal response, X1 = Phenylalanine ammonia lyase, X2 = Polyphenol oxidase, X3 = Total phenols, X4 = Flavonols, X5 = Free amino acids. | |
| [1] | Bacheller J D, Romeo J T.1992. Biotic and abiotic stress effects on nitrogen chemistry in the salt marsh cordgrass Spartina alterniflora(Poaceae). Chemoecology, 3: 74-80. |
| [2] | Balachiranjeevi C H, Prahalada G D, Mahender A, Jamaloddin M, Sevilla M A L, Marfori-Nazarea C M, Vinarao R, Sushanto U, Baehaki S E, Li Z K, Ali J.2019. Identification of a novel locus, Bph38(t), conferring resistance to brown planthopper, 215(11): 185. |
| [3] | Bhanu K V, Lakshmi V J, Katti G, Reddy A V.2014. Antibiosis and tolerance mechanisms of resistance in rice varieties carrying brown planthopper resistance genes. Asian J Biol Life Sci, 3: 108-113. |
| [4] | Bhonwong A, Stout M J, Attajarusit J, Tantasawat P.2009. Defensive role of tomato polyphenol oxidases against cotton bollworm (Helicoverpa armigera) and beet armyworm(Spodoptera exigua). J Chem Ecol, 35(1): 28-38. |
| [5] | Catindig J L A, Arida G S, Baehaki S E, Bentur J S, Cuong L Q, Norowi M, Rattanakarn W, Sriratanasak W, Xia J, Lu Z.2009. Situation of planthoppers in Asia. In: Heong K L, Hardy B. Planthoppers: New Threats to the Sustainability of Intensive Rice Production Systems in Asia. Los Banos, the Philippines: 191-220. |
| [6] | Dixit G, Praveen A, Tripathi T, Yadav V K, Verma P C.2017. Herbivore-responsive cotton phenolics and their impact on insect performance and biochemistry. J Asia-Pac Entomol, 20(2): 341-351. |
| [7] | Du B, Zhang W L, Liu B F, Hu J, Wei Z, Shi Z Y, He R F, Zhu L L, Chen R Z, Han B, He G C.2009. Identification and characterization of Bph14, a gene conferring resistance to brown planthopper in rice. Proc Natl Acad Sci USA, 106: 22163-22168. |
| [8] | Duan C X, Yu J J, Bai J Y, Zhu Z D, Wang X M.2014. Induced defence responses in rice plants against small brown planthopper infestation. Crop J, 2(1): 55-62. |
| [9] | Felton G W, Bi J L, Summers C B, Mueller A J, Duffey S S.1994. Potential role of lipoxygenases in defence against insect herbivory. J Chem Ecol, 20: 651-666. |
| [10] | Fischer M K, Shingleton A W.2001. Host plant and ants influence the honeydew sugar composition of aphids. Funct Ecol, 15(4): 544-550. |
| [11] | Fujita D, Kohli A, Horgan F G.2013. Rice resistance to planthoppers and leafhoppers. Crit Rev Plant Sci, 32(3): 162-191. |
| [12] | Gangaraju P, Shivashankar T, Lohithaswa H C.2017. Genetic basis of resistance to brown planthopper (Nilaparvata lugens Stål) in local landraces of rice. Int J Curr Microbiol App Sci, 6: 3388-3393. |
| [13] | Han Y Q, Wu C, Yang L, Zhang D Y, Xiao Y T.2018. Resistance to Nilaparvata lugens in rice lines introgressed with the resistance genes Bph14 and Bph15 and related resistance types. PLoS One, 13(6): e0198630. |
| [14] | Hao P Y, Liu C X, Wang Y Y, Chen R Z, Tang M, Du B, Zhu L L, He G C.2008. Herbivore induced callose deposition on the sieve plates of rice: An important mechanism for host resistance. Plant Physiol, 146(4): 1810-1820. |
| [15] | He J P, Chen F D, Chen S M, Lv G S, Deng Y M, Fang W M, Liu Z L, Guan Z Y, He C Y.2011. Chrysanthemum leaf epidermal surface morphology and antioxidant and defence enzyme activity in response to aphid infestation. J Plant Physiol, 168(7): 687-693. |
| [16] | Howe G A, Jander G.2008. Plant immunity to insect herbivores. Ann Rev Plant Biol, 59: 41-66. |
| [17] | Huang Z, He G, Shu L, Li X, Zhang Q.2001. Identification and mapping of two brown planthopper resistance genes in rice. Theor Appl Genet, 102: 929-934. |
| [18] | Jayasimha G T, Nalini R, Chinniah C, Muthamilan M, Mini M L.2015. Evaluation of biochemical constituents in healthy and brown planthopper,Nilaparvata lugens(Stål)(Hemiptera: Delphacidae) damaged rice plants. Curr Biot, 9: 129-136. |
| [19] | Kaur H, Salh P K, Singh B.2017. Role of defence enzymes and phenolics in resistance of wheat crop (Triticum aestivum L.) towards aphid complex. J Plant Interact, 12: 304-311. |
| [20] | Khush G S, Brar D S.1991. Genetics of resistance to insects in crop plants. Adv Agron, 45: 223-274. |
| [21] | Kumar K, Sarao P S, Bhatia D, Neelam K, Kaur A, Mangat G S, Brar D S, Singh K.2018. High-resolution genetic mapping of a novel brown planthopper resistance locus,Bph34 in Oryza sativa L. × Oryza nivara(Sharma & Shastry) derived interspecific F2 population. Theor Appl Genet, 131(5): 1163-1171. |
| [22] | Li J, Chen Q H, Wang L Q, Liu J, Shang K K, Hua H X.2011. Biological effects of rice harbouring Bph14 and Bph15 on brown planthopper, Nilaparvata lugens. Pest Manag Sci, 67(5): 528-534. |
| [23] | Nanda U K, Dash D, Rath L K.2000. Biochemical basis of resistance in rice to the brown planthopper Nilaparvata lugens. Ind J Entomol, 62: 239-241. |
| [24] | Rani P U, Jyothsna Y.2010. Biochemical and enzymatic changes in rice as a mechanism of defence. Acta Physiol Plant, 32: 695-701. |
| [25] | Rani P U, Ravibabu M V.2011. Allelochemicals in castor (Ricinus communis) plants and their impact on pest larval feeding as anti-herbivore defensive. Allel J, 27(2): 263-276. |
| [26] | Rani P U, Pratyusha S.2014. Role of castor plant phenolics on performance of its two herbivores and their impact on egg parasitoid behaviour. BioControl, 59: 513-524. |
| [27] | Sarao P S, Sahi G K, Kumari N, Mangat G S, Patra B C, Singh K.2016. Donors for resistance to brown planthopper Nilaparvata lugens(Stål) from wild rice species. Rice Sci, 23(4): 219-224. |
| [28] | Scott M I, Thaler S J, Scott G F.2010. Response of a generalist herbivore Trichoplusia ni to jasmonate-mediated induced defence in tomato. J Chem Ecol, 36: 490-499. |
| [29] | Seino Y, Suzuki Y, Sogawa K.1996. An ovicidal substance produced by rice plants in response to oviposition by the whitebacked planthopper,Sogatella furcifera(HORVATH) (Homoptera: Delphacidae). Appl Entomol Zool, 31(4): 467-473. |
| [30] | Sogawa K.1982. The rice brown planthopper: Feeding physiology and host plant interactions. Ann Rev Entomol, 27: 49-73. |
| [31] | Sogawa K, Liu G J, Zhu C G.2003. Inheritance of whitebacked planthopper resistance in Chinese japonica rice Chunjiang 06. Chin J Rice Sci, 17: 73-76. (in Chinese with English abstract) |
| [32] | Taggar G K, Gill R S, Gupta A K, Singh S.2014. Induced changes in the antioxidative compounds of Vigna mungo genotypes due to infestation by Bemisia tabaci(Gennadius). J Environ Biol, 35(6): 1037-1045. |
| [33] | Tallamy D W, Stull J, Ehresman N P, Gorski P M, Mason C E.1997. Cucurbitacins as feeding and oviposition deterrents to insects. Environ Entomol, 26(3): 678-683. |
| [34] | Vanitha K, Suresh S, Gunathilagaraj K.2011. Influence of brown planthopper, Nilaparvava lugens(Stål) feeding on nutritional biochemistry of rice plant. Oryza, 48: 142-146. |
| [35] | War A R, Paulraj M G, War M Y, Ignacimuthu S.2012. Herbivore induced resistance in different groundnut germplasm lines to Asian armyworm, Spodoptera litura (Fab.) (Lepidoptera: Noctuidae). Acta Physiol Plant, 34: 343-352. |
| [36] | War A R, Paulraj M G, Ignacimuthu S, Sharma H C.2013. Defensive responses in groundnut against chewing and sap- sucking insects. J Plant Growth Regul, 32: 259-272. |
| [37] | War A R, Paulraj M G, Ignacimuthu S, Sharma H C.2015. Induced resistance to Helicoverpa armigera through exogenous application of jasmonic acid and salicylic acid in groundnut, Arachis hypogaea. Pest Manag Sci, 71(1): 72-82. |
| [38] | War A R, Taggar G K, Hussain B, Taggar M S, Nair R M, Sharma H C.2018. Plant defence against herbivory and insect adaptations. AoB Plants, 10: ply037. |
| [39] | Win S S, Muhamad R, Ahmad Z A M, Adam N A.2011. Population fluctuations of brown planthopper Nilaparvata lugens Stal. and whitebacked planthopper Sogatella furcifera Horvath on rice. J Entomol, 8(2): 183-190. |
| [40] | Yamasaki M, Yoshimura A, Yasui H.2003. Genetic basis of ovicidal response to whitebacked planthopper (Sogatella furcifera Horvath) in rice(Oryza sativa L.). Mol Breeding, 12: 133-143. |
| [41] | Yang Y L, Xu J, Leng Y J, Xiong G S, Hu J, Zhang G H, Huang L C, Wang L, Guo L B, Li J Y, Chen F, Qian Q, Zeng D L.2014. Quantitative trait loci identification, fine mapping and gene expression profiling for ovicidal response to whitebacked planthopper (Sogatella furcifera Horvath) in rice(Oryza sativa L.). BMC Plant Biol, 14: 145. |
| [42] | Yang L, Han Y Q, Li P, Wen L Z, Hou M L.2017. Silicon amendment to rice plants impairs sucking behaviors and population growth in the phloem feeder Nilaparvata lugens(Hemiptera: Delphacidae). Sci Rep, 7: 1101. |
| [43] | Zhao L Y, Chen J L, Cheng D F, Sun J R, Liu Y, Tian Z.2009. Biochemical and molecular characterizations of Sitobion avenae-induced wheat defence responses. Crop Prot, 28(5): 435-442. |
| [44] | Zhou S, Lou Y R, Tzin V, Jander G.2015. Alternation of plant primary metabolism in response to insect herbivory. Plant Physiol, 169(3): 1488-1498. |
| No related articles found! |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||