
Rice Science ›› 2019, Vol. 26 ›› Issue (4): 220-228.DOI: 10.1016/j.rsci.2019.04.004
收稿日期:2019-02-20
接受日期:2019-04-26
出版日期:2019-07-28
发布日期:2019-04-04
. [J]. Rice Science, 2019, 26(4): 220-228.
| Year | Population | Segregating region | Number of lines | |||||
| Generation | Name | Marker | Physical position (bp) | NILZS97 | NILMY46 | |||
| 2014 | BC2F12 | W14-1 | RM212-Wn33252 | 33 053 493-33 252 459 | 20 | 20 | ||
| W14-2 | RM212-Wn33252 | 33 053 493-33 252 459 | 20 | 19 | ||||
| W14-3 | Wn33252-Wn33304 | 33 252 459-33 304 754 | 40 | 40 | ||||
| 2016 | BC2F14 | W16-1 | Wn33011-Wn33164 | 33 011 637-33 164 422 | 19 | 14 | ||
| W16-2 | Wn33186-Wn33252 | 33 186 760-33 252 459 | 33 | 33 | ||||
| 2017 | BC2F16 | W17-1 | Wn33011-Wn33186 | 33 011 637-33 186 760 | 33 | 34 | ||
| W17-2 | RM212-Wn33252 | 33 053 493-33 252 459 | 33 | 34 | ||||
| W17-3 | Wn33186-Wn33252 | 33 186 760-33 252 459 | 34 | 35 | ||||
| 2018 | BC2F17 | W18-1 | Wn32997-Wn33011 | 32 996 766-33 011 637 | 30 | 30 | ||
| W18-2 | RM212-Wn33186 | 33 053 493-33 186 760 | 28 | 28 | ||||
| W18-3 | Wn33089-Wn33252 | 33 089 090-33 252 459 | 29 | 30 | ||||
| NILZS97 and NILMY46 are near isogenic lines (NILs) having Zhenshan 97 and Milyang 46 homozygous genotypes in the segregating regions, respectively. | ||||||||
Table 1 Eleven rice populations used for QTL analysis in this study.
| Year | Population | Segregating region | Number of lines | |||||
| Generation | Name | Marker | Physical position (bp) | NILZS97 | NILMY46 | |||
| 2014 | BC2F12 | W14-1 | RM212-Wn33252 | 33 053 493-33 252 459 | 20 | 20 | ||
| W14-2 | RM212-Wn33252 | 33 053 493-33 252 459 | 20 | 19 | ||||
| W14-3 | Wn33252-Wn33304 | 33 252 459-33 304 754 | 40 | 40 | ||||
| 2016 | BC2F14 | W16-1 | Wn33011-Wn33164 | 33 011 637-33 164 422 | 19 | 14 | ||
| W16-2 | Wn33186-Wn33252 | 33 186 760-33 252 459 | 33 | 33 | ||||
| 2017 | BC2F16 | W17-1 | Wn33011-Wn33186 | 33 011 637-33 186 760 | 33 | 34 | ||
| W17-2 | RM212-Wn33252 | 33 053 493-33 252 459 | 33 | 34 | ||||
| W17-3 | Wn33186-Wn33252 | 33 186 760-33 252 459 | 34 | 35 | ||||
| 2018 | BC2F17 | W18-1 | Wn32997-Wn33011 | 32 996 766-33 011 637 | 30 | 30 | ||
| W18-2 | RM212-Wn33186 | 33 053 493-33 186 760 | 28 | 28 | ||||
| W18-3 | Wn33089-Wn33252 | 33 089 090-33 252 459 | 29 | 30 | ||||
| NILZS97 and NILMY46 are near isogenic lines (NILs) having Zhenshan 97 and Milyang 46 homozygous genotypes in the segregating regions, respectively. | ||||||||
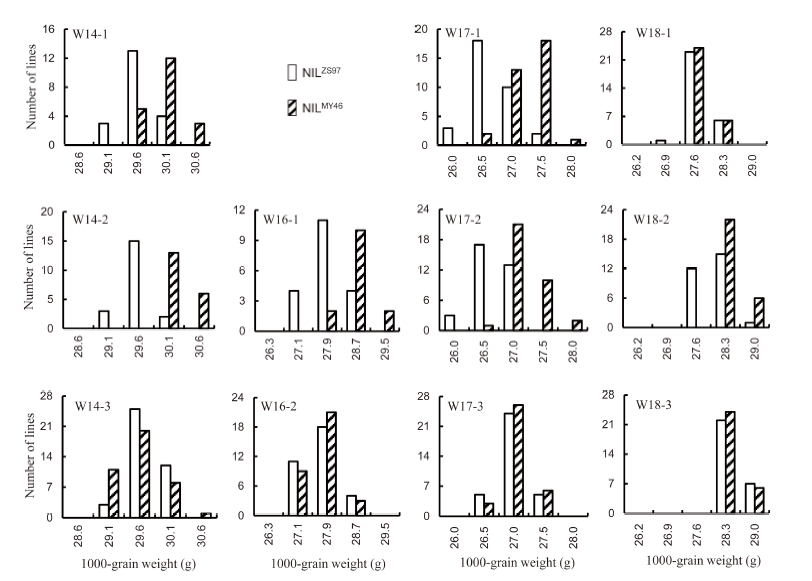
Fig. 1. Distribution of 1000-grain weight in 11 near isogenic line populations of rice.NILZS97 and NILMY46 are near isogenic lines (NILs) having Zhenshan 97 and Milyang 46 homozygous genotypes in the segregating regions, respectively.
| Marker | Forward primer (5'-3') | Reverse primer (5'-3') |
|---|---|---|
| Wn32886 | CGTGTTACTAACCCAGTCAACTCAGG | CAATAGTAGCAGCTAAATTGGCGTTC |
| Wn32922 | AAGAAGAACATTTGCTCACAG | TGCAGTCTATTGTTGACCA |
| Wn32997 | GTCAACCCAAATGTAGTAAGC | CCGCCGAGGTGAAGTTGTAG |
| Wn33011 | TACAATCTACTATCCCGGACT | CTCGCAATTTACATGCAAACT |
| Wn33072 | CTGGACAACATCCCCATGCT | GACCGGAAACCAAATACACCC |
| Wn33084 | TTGTGACCGCATTAATAAGCC | TAATATTTTATGCGTAACTTG |
| Wn33089 | GCTGCCGTGCAATTCGTAGTT | GAGGGATCGCGATGGGGGAGT |
| Wn33164 | TACGCACTTGCCCTTAGATGTC | TTTCCTATCTATTTATTGCCCTA |
| Wn33186 | AGTTAGTGCAGCAATCTACGTCCT | AGCTCCTCCAGTGACGATACGG |
| Wn33252 | GCATGTATCAAAGATTCGATGAGA | TGAAAACTCATGGCTACGCT |
| Wn33268 | CGTATATGCTCGTGACTCTG | ATCAGCGAGACAACGACTTCC |
| Wn33304 | ATCCAAAATAGTTGAGGGCAT | TTATTTAGCTAGATTATTATAGGCTGG |
Supplemental Table 1 InDel markers used in this study.
| Marker | Forward primer (5'-3') | Reverse primer (5'-3') |
|---|---|---|
| Wn32886 | CGTGTTACTAACCCAGTCAACTCAGG | CAATAGTAGCAGCTAAATTGGCGTTC |
| Wn32922 | AAGAAGAACATTTGCTCACAG | TGCAGTCTATTGTTGACCA |
| Wn32997 | GTCAACCCAAATGTAGTAAGC | CCGCCGAGGTGAAGTTGTAG |
| Wn33011 | TACAATCTACTATCCCGGACT | CTCGCAATTTACATGCAAACT |
| Wn33072 | CTGGACAACATCCCCATGCT | GACCGGAAACCAAATACACCC |
| Wn33084 | TTGTGACCGCATTAATAAGCC | TAATATTTTATGCGTAACTTG |
| Wn33089 | GCTGCCGTGCAATTCGTAGTT | GAGGGATCGCGATGGGGGAGT |
| Wn33164 | TACGCACTTGCCCTTAGATGTC | TTTCCTATCTATTTATTGCCCTA |
| Wn33186 | AGTTAGTGCAGCAATCTACGTCCT | AGCTCCTCCAGTGACGATACGG |
| Wn33252 | GCATGTATCAAAGATTCGATGAGA | TGAAAACTCATGGCTACGCT |
| Wn33268 | CGTATATGCTCGTGACTCTG | ATCAGCGAGACAACGACTTCC |
| Wn33304 | ATCCAAAATAGTTGAGGGCAT | TTATTTAGCTAGATTATTATAGGCTGG |
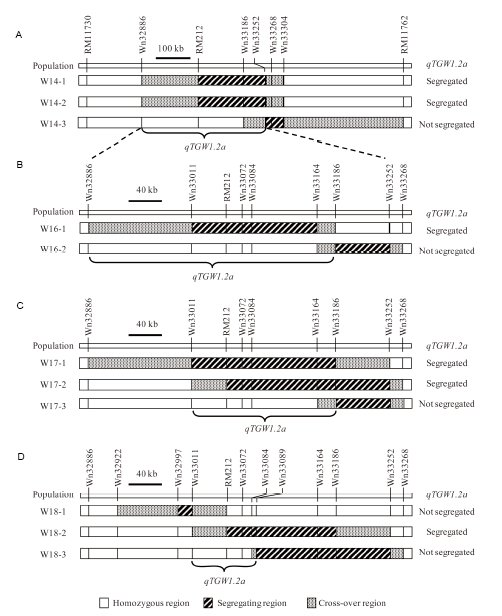
Fig. 2. Genotypic compositions of the near isogenic line populations in the target region.A, Three BC2F12 populations. B, Two BC2F14 populations. C, Three BC2F16 populations. D, Three BC2F17 populations.
Supplemental Fig. 2. Distribution of grain length and grain width in 11 near isogenic line populations of rice.A. Three populations in BC2F12. B. Two populations in BC2F14. C. Three populations in BC2F16. D. Three populations in BC2F17.
| Generation | Name | Trait | Phenotype (Mean ± SD) | P value | A | R2 (%) | |
| NILZS97 | NILMY46 | ||||||
| BC2F12 | W14-1 | TGW (g) | 29.30 ± 0.27 | 29.77 ± 0.30 | <0.0001 | 0.23 | 26.15 |
| GL (mm) | 8.515 ± 0.040 | 8.519 ± 0.041 | 0.7831 | ||||
| GW (mm) | 3.278 ± 0.016 | 3.304 ± 0.020 | <0.0001 | 0.013 | 21.60 | ||
| W14-2 | TGW (g) | 29.27 ± 0.27 | 29.99 ± 0.27 | <0.0001 | 0.36 | 41.65 | |
| GL (mm) | 8.561 ± 0.042 | 8.576 ± 0.034 | 0.2397 | ||||
| GW (mm) | 3.303 ± 0.019 | 3.355 ± 0.019 | <0.0001 | 0.026 | 42.38 | ||
| W14-3 | TGW (g) | 29.44 ± 0.28 | 29.33 ± 0.34 | 0.1240 | |||
| GL (mm) | 8.545 ± 0.051 | 8.557 ± 0.049 | 0.3004 | ||||
| GW (mm) | 3.242 ± 0.021 | 3.226 ± 0.026 | 0.0038 | -0.008 | 6.00 | ||
| BC2F14 | W16-1 | TGW (g) | 27.51 ± 0.43 | 28.28 ± 0.37 | <0.0001 | 0.38 | 31.23 |
| GL (mm) | 8.261 ± 0.048 | 8.319 ± 0.041 | 0.0013 | 0.029 | 20.34 | ||
| GW (mm) | 3.079 ± 0.017 | 3.111 ± 0.016 | <0.0001 | 0.016 | 32.37 | ||
| W16-2 | TGW (g) | 27.30 ± 0.46 | 27.32 ± 0.39 | 0.8195 | |||
| GL (mm) | 8.302 ± 0.043 | 8.291 ± 0.045 | 0.3423 | ||||
| GW (mm) | 3.064 ± 0.021 | 3.065 ± 0.024 | 0.8812 | ||||
| BC2F16 | W17-1 | TGW (g) | 26.40 ± 0.32 | 26.99 ± 0.31 | <0.0001 | 0.29 | 35.39 |
| GL (mm) | 8.207 ± 0.042 | 8.246 ± 0.038 | 0.0002 | 0.019 | 11.55 | ||
| GW (mm) | 3.028 ± 0.019 | 3.070 ± 0.025 | <0.0001 | 0.021 | 35.80 | ||
| W17-2 | TGW (g) | 26.38 ± 0.26 | 26.94 ± 0.29 | <0.0001 | 0.28 | 33.59 | |
| GL (mm) | 8.195 ± 0.037 | 8.263 ± 0.036 | <0.0001 | 0.034 | 28.15 | ||
| GW (mm) | 3.042 ± 0.019 | 3.064 ± 0.021 | <0.0001 | 0.011 | 16.21 | ||
| W17-3 | TGW (g) | 26.77 ± 0.23 | 26.78 ± 0.23 | 0.8769 | |||
| GL (mm) | 8.306 ± 0.027 | 8.302 ± 0.031 | 0.5432 | ||||
| GW (mm) | 3.031 ± 0.016 | 3.031 ± 0.014 | 0.9496 | ||||
| BC2F17 | W18-1 | TGW (g) | 27.44 ± 0.21 | 27.46 ± 0.18 | 0.7023 | ||
| GL (mm) | 8.323 ± 0.023 | 8.315 ± 0.025 | 0.2390 | ||||
| GW (mm) | 3.146 ± 0.027 | 3.149 ± 0.023 | 0.5819 | ||||
| W18-2 | TGW (g) | 27.64 ± 0.34 | 28.17 ± 0.24 | <0.0001 | 0.26 | 36.65 | |
| GL (mm) | 8.340 ± 0.036 | 8.352 ± 0.025 | 0.1796 | ||||
| GW (mm) | 3.140 ± 0.031 | 3.182 ± 0.025 | <0.0001 | 0.021 | 25.49 | ||
| W18-3 | TGW (g) | 28.15 ± 0.18 | 28.19 ± 0.13 | 0.3866 | |||
| GL (mm) | 8.373 ± 0.027 | 8.367 ± 0.023 | 0.4148 | ||||
| GW (mm) | 3.170 ± 0.016 | 3.169 ± 0.016 | 0.8698 | ||||
| TGW, 1000-grain weight; GL, Grain length; GW, Grain width; A, Additive effect of replacing a Zhenshan 97 allele with a Milyang 46 allele; R2, Proportion of phenotypic variance explained by the QTL effect. | |||||||
Table 3 Phenotypic difference between the two genotypic groups in each population.
| Generation | Name | Trait | Phenotype (Mean ± SD) | P value | A | R2 (%) | |
| NILZS97 | NILMY46 | ||||||
| BC2F12 | W14-1 | TGW (g) | 29.30 ± 0.27 | 29.77 ± 0.30 | <0.0001 | 0.23 | 26.15 |
| GL (mm) | 8.515 ± 0.040 | 8.519 ± 0.041 | 0.7831 | ||||
| GW (mm) | 3.278 ± 0.016 | 3.304 ± 0.020 | <0.0001 | 0.013 | 21.60 | ||
| W14-2 | TGW (g) | 29.27 ± 0.27 | 29.99 ± 0.27 | <0.0001 | 0.36 | 41.65 | |
| GL (mm) | 8.561 ± 0.042 | 8.576 ± 0.034 | 0.2397 | ||||
| GW (mm) | 3.303 ± 0.019 | 3.355 ± 0.019 | <0.0001 | 0.026 | 42.38 | ||
| W14-3 | TGW (g) | 29.44 ± 0.28 | 29.33 ± 0.34 | 0.1240 | |||
| GL (mm) | 8.545 ± 0.051 | 8.557 ± 0.049 | 0.3004 | ||||
| GW (mm) | 3.242 ± 0.021 | 3.226 ± 0.026 | 0.0038 | -0.008 | 6.00 | ||
| BC2F14 | W16-1 | TGW (g) | 27.51 ± 0.43 | 28.28 ± 0.37 | <0.0001 | 0.38 | 31.23 |
| GL (mm) | 8.261 ± 0.048 | 8.319 ± 0.041 | 0.0013 | 0.029 | 20.34 | ||
| GW (mm) | 3.079 ± 0.017 | 3.111 ± 0.016 | <0.0001 | 0.016 | 32.37 | ||
| W16-2 | TGW (g) | 27.30 ± 0.46 | 27.32 ± 0.39 | 0.8195 | |||
| GL (mm) | 8.302 ± 0.043 | 8.291 ± 0.045 | 0.3423 | ||||
| GW (mm) | 3.064 ± 0.021 | 3.065 ± 0.024 | 0.8812 | ||||
| BC2F16 | W17-1 | TGW (g) | 26.40 ± 0.32 | 26.99 ± 0.31 | <0.0001 | 0.29 | 35.39 |
| GL (mm) | 8.207 ± 0.042 | 8.246 ± 0.038 | 0.0002 | 0.019 | 11.55 | ||
| GW (mm) | 3.028 ± 0.019 | 3.070 ± 0.025 | <0.0001 | 0.021 | 35.80 | ||
| W17-2 | TGW (g) | 26.38 ± 0.26 | 26.94 ± 0.29 | <0.0001 | 0.28 | 33.59 | |
| GL (mm) | 8.195 ± 0.037 | 8.263 ± 0.036 | <0.0001 | 0.034 | 28.15 | ||
| GW (mm) | 3.042 ± 0.019 | 3.064 ± 0.021 | <0.0001 | 0.011 | 16.21 | ||
| W17-3 | TGW (g) | 26.77 ± 0.23 | 26.78 ± 0.23 | 0.8769 | |||
| GL (mm) | 8.306 ± 0.027 | 8.302 ± 0.031 | 0.5432 | ||||
| GW (mm) | 3.031 ± 0.016 | 3.031 ± 0.014 | 0.9496 | ||||
| BC2F17 | W18-1 | TGW (g) | 27.44 ± 0.21 | 27.46 ± 0.18 | 0.7023 | ||
| GL (mm) | 8.323 ± 0.023 | 8.315 ± 0.025 | 0.2390 | ||||
| GW (mm) | 3.146 ± 0.027 | 3.149 ± 0.023 | 0.5819 | ||||
| W18-2 | TGW (g) | 27.64 ± 0.34 | 28.17 ± 0.24 | <0.0001 | 0.26 | 36.65 | |
| GL (mm) | 8.340 ± 0.036 | 8.352 ± 0.025 | 0.1796 | ||||
| GW (mm) | 3.140 ± 0.031 | 3.182 ± 0.025 | <0.0001 | 0.021 | 25.49 | ||
| W18-3 | TGW (g) | 28.15 ± 0.18 | 28.19 ± 0.13 | 0.3866 | |||
| GL (mm) | 8.373 ± 0.027 | 8.367 ± 0.023 | 0.4148 | ||||
| GW (mm) | 3.170 ± 0.016 | 3.169 ± 0.016 | 0.8698 | ||||
| TGW, 1000-grain weight; GL, Grain length; GW, Grain width; A, Additive effect of replacing a Zhenshan 97 allele with a Milyang 46 allele; R2, Proportion of phenotypic variance explained by the QTL effect. | |||||||
| Locus name | Gene product name |
|---|---|
| LOC_Os01g57120 | Transposon protein, putative, unclassified |
| LOC_Os01g57130 | Retrotransposon protein, putative, Ty3-gypsy subclass, expressed |
| LOC_Os01g57140 | Hypothetical protein |
| LOC_Os01g57150 | SR protein related family member, putative, expressed |
| LOC_Os01g57160 | Transposon protein, putative, CACTA, En/Spm sub-class, expressed |
| LOC_Os01g57170 | Expressed protein, uncharacterized |
| LOC_Os01g57180 | Hypothetical protein |
| LOC_Os01g57190 | Transposon protein, putative, unclassified, expressed |
| LOC_Os01g57210 | Katanin p80 WD40 repeat-containing subunit B1 homolog 1, putative, expressed |
| LOC_Os01g57220 | Secretory carrier-associated membrane protein, putative, expressed |
| LOC_Os01g57230 | BTBN1 - Bric-a-Brac, Tramtrack, Broad Complex BTB domain with non-phototropic hypocotyl 3 NPH3 and coiled-coil domains, expressed |
| LOC_Os01g57240 | Expressed protein, uncharacterized |
| LOC_Os01g57250 | Expressed protein, uncharacterized |
Supplemental Table 2 Supplemental Table 2. Annotated genes in the 77.5-kb region for qTGW1.2a.
| Locus name | Gene product name |
|---|---|
| LOC_Os01g57120 | Transposon protein, putative, unclassified |
| LOC_Os01g57130 | Retrotransposon protein, putative, Ty3-gypsy subclass, expressed |
| LOC_Os01g57140 | Hypothetical protein |
| LOC_Os01g57150 | SR protein related family member, putative, expressed |
| LOC_Os01g57160 | Transposon protein, putative, CACTA, En/Spm sub-class, expressed |
| LOC_Os01g57170 | Expressed protein, uncharacterized |
| LOC_Os01g57180 | Hypothetical protein |
| LOC_Os01g57190 | Transposon protein, putative, unclassified, expressed |
| LOC_Os01g57210 | Katanin p80 WD40 repeat-containing subunit B1 homolog 1, putative, expressed |
| LOC_Os01g57220 | Secretory carrier-associated membrane protein, putative, expressed |
| LOC_Os01g57230 | BTBN1 - Bric-a-Brac, Tramtrack, Broad Complex BTB domain with non-phototropic hypocotyl 3 NPH3 and coiled-coil domains, expressed |
| LOC_Os01g57240 | Expressed protein, uncharacterized |
| LOC_Os01g57250 | Expressed protein, uncharacterized |
| [1] | Bai X F, Wu B, Xing Y Z.2012. Yield-related QTLs and their applications in rice genetic improvement.J Integr Plant Biol, 54(5): 300-311. |
| [2] | Chen J Y, Zhang H W, Zhang H L, Ying J Z, Ma L Y, Zhuang J Y.2018. Natural variation atqHd1 affects heading date acceleration at high temperatures with pleiotropism for yield traits in rice. BMC Plant Biol, 18: 112. |
| [3] | Chen X, Temnykh S, Xu Y, Cho Y G, McCouch S R.1997. Development of a microsatellite framework map providing genome-wide coverage in rice (Oryza sativa L.). Theor Appl Genet, 95(4): 553-567. |
| [4] | Dai W M, Zhang K Q, Wu J R, Wang L, Duan B W, Zheng K L, Cai R, Zhuang J Y.2008. Validating a segment on the short arm of chromosome 6 responsible for genetic variation in the hull silicon content and yield traits of rice.Euphytica, 160(3): 317-324. |
| [5] | Dong Q, Zhang Z H, Wang L L, Zhu Y J, Fan Y Y, Mou T M, Ma L Y, Zhuang J Y.2018. Dissection and fine-mapping of two QTL for grain size linked in a 460-kb region on chromosome 1 of rice. Rice, 11(1): 44. |
| [6] | Duan P G, Ni S, Wang J M, Zhang B L, Xu R, Wang Y X, Chen H Q, Zhu X D, Li Y H.2015. Regulation ofOsGRF4 by OsmiR396 controls grain size and yield in rice. Nat Plants, 2: 15203. |
| [7] | Duan P G, Xu J S, Zeng D L, Zhang B L, Geng M F, Zhang G Z, Huang K, Huang L J, Xu R, Ge S, Qian Q, Li Y H.2017. Natural variation in the promoter ofGSE5 contributes to grain size diversity in rice. Mol Plant, 10(5): 685-694. |
| [8] | Fan C C, Xing Y Z, Mao H L, Lu T T, Han B, Xu C G, Li X H, Zhang Q F.2006. GS3, a major QTL for grain length and weight and minor QTL for grain width and thickness in rice, encodes a putative transmembrane protein. Theor Appl Genet, 112(6): 1164-1171. |
| [9] | Hu J, Wang Y X, Fang Y X, Zeng L J, Xu J, Yu H P, Shi Z Y, Pan J J, Zhang D, Kang S J, Zhu L, Dong G J, Guo L B, Zeng D L, Zhang G H, Xie L H, Xiong G S, Li J Y, Qian Q.2015. A rare allele ofGS2 enhances grain size and grain yield in rice. Mol Plant, 8(10): 1455-1465. |
| [10] | Hu Z J, Lu S J, Wang M J, He H H, Sun L, Wang H R, Liu X H, Jiang L, Sun J L, Xin X Y, Kong W, Chu C C, Xue H W, Yang J S, Luo X J, Liu J X.2018. A novel QTL qTGW3 encodes the GSK3/SHAGGY-like kinase OsGSK5/OsSK41 that interacts with OsARF4 to negatively regulate grain size and weight in rice. Mol Plant, 11(5): 736-749. |
| [11] | Huang R Y, Jiang L R, Zheng J S, Wang T S, Wang H C, Huang Y M, Hong Z L.2013. Genetic bases of rice grain shape: So many genes, so little known.Trends Plant Sci, 18(4): 218-226. |
| [12] | Ishimaru K, Hirotsu N, Madoka Y, Murakami N, Hara N, Onodera H, Kashiwagi T, Ujiie K, Shimizu B, Onishi A, Miyagawa H, Katoh E.2013. Loss of function of the IAA-glucose hydrolase geneTGW6 enhances rice grain weight and increases yield. Nat Genet, 45(6): 707-711. |
| [13] | Li N, Xu R, Duan P G, Li Y H.2018. Control of grain size in rice.Plant Reprod, 31(3): 237-251. |
| [14] | Li Y B, Fan C C, Xing Y Z, Jiang Y H, Luo L J, Sun L, Shao D, Xu C J, Li X H, Xiao J H, He Y Q, Zhang Q F.2011. Natural variation in GS5 plays an important role in regulating grain size and yield in rice. Nat Genet, 43(12): 1266-1269. |
| [15] | Liu J F, Chen J, Zheng X M, Wu F Q, Lin Q B, Heng Y Q, Tian P, Cheng Z J, Yu X W, Zhou K N, Zhang X, Guo X P, Wang J L, Wang H Y, Wan J M.2017. GW5 acts in the brassinosteroid signalling pathway to regulate grain width and weight in rice. Nat Plants, 3: 17043. |
| [16] | Liu Q, Han R X, Wu K, Zhang J Q, Ye Y F, Wang S S, Chen J F, Pan Y J, Li Q, Xu X P, Zhou J W, Tao D Y, Wu Y J, Fu X D.2018. G-protein βγ subunits determine grain size through interaction with MADS-domain transcription factors in rice.Nat Commun, 9: 852. |
| [17] | Mackay T F C, Stone E A, Ayroles J F.2009. The genetics of quantitative traits: Challenges and prospects.Nat Rev Genet, 10(8): 565-577. |
| [18] | Matsubara K, Ogiso-Tanaka E, Hori K, Ebana K, Ando T, Yano M.2012. Natural variation inHd17, a homolog of Arabidopsis ELF3 that is involved in rice photoperiodic flowering. Plant Cell Physiol, 53(4): 709-716. |
| [19] | Qi L, Ding Y B, Zheng X M, Xu R, Zhang L Z, Wang Y Y, Wang X N, Zhang L F, Cheng Y L, Yang Q W.2018. Fine mapping and identification of a novel locusqGL12.2 control grain length in wild rice(Oryza rufipogon Griff.). Theor Appl Genet, 131(7): 1497-1508. |
| [20] | Qi P, Lin Y S, Song X J, Shen J B, Huang W, Shan J X, Zhu M Z, Jiang L, Gao J P, Lin H X.2012. The novel quantitative trait locusGL3.1 controls rice grain size and yield by regulating Cyclin-T1;3. Cell Res, 22(12): 1666-1680. |
| [21] | SAS Institute Inc.1999. SAS/STAT User’s Guide . Cary:SAS Institute. |
| [22] | Shao G N, Tang S Q, Luo J, Jiao G A, Wei X J, Tang A, Wu J L, Zhuang J Y, Hu P S.2010. Mapping ofqGL7-2, a grain length QTL on chromosome 7 of rice. J Genet Genom, 37(8): 523-531. |
| [23] | Si L Z, Chen J Y, Huang X H, Gong H, Luo J H, Hou Q Q, Zhou T Y, Lu T T, Zhu J J, Shangguan Y Y, Chen E W, Gong G X, Zhao Q, Jing Y F, Zhao Y, Li Y, Cui L L, Fan D L, Lu Y Q, Weng Q J, Wang Y C, Zhan Q L, Liu K Y, Wei X H, An K, An G, Han B.2016. OsSPL13 controls grain size in cultivated rice. Nat Genet, 48: 447-456. |
| [24] | Song X J, Huang W, Shi M, Zhu M Z, Lin H X.2007. A QTL for rice grain width and weight encodes a previously unknown RING-type E3 ubiquitin ligase.Nat Genet, 39: 623-630. |
| [25] | Song X J, Kuroha T, Ayano M, Furuta T, Nagai K, Komeda N, Segami S, Miura K, Ogawa D, Kamura T, Suzuki T, Higashiyama T, Yamasaki M, Mori H, Inukai Y, Wu J Z, Kitano H, Sakakibara H, Jacobsen S E, Ashikari M.2015. Rare allele of a previously unidentified histone H4 acetyltransferase enhances grain weight, yield, and plant biomass in rice.Proc Natl Acad Sci USA, 112(1): 76-81. |
| [26] | Wang L L, Chen Y Y, Guo L, Zhang H W, Fan Y Y, Zhuang J Y.2015. Dissection ofqTGW1.2 to three QTLs for grain weight and grain size in rice(Oryza sativa L.). Euphytica, 202(1): 119-127. |
| [27] | Wang S K, Wu K, Yuan Q B, Liu X Y, Liu Z B, Lin X, Y Zeng R Z, Zhu H T, Dong G J, Qian Q, Zhang G Q, Fu X D.2012. Control of grain size, shape and quality byOsSPL16 in rice. Nat Genet, 44(8): 950-954. |
| [28] | Wang S K, Li S, Liu Q, Wu K, Zhang J Q, Wang S S, Wang Y, Chen X B, Zhang Y, Gao C X, Wang F, Huang H X, Fu X D.2015. TheOsSPL16-GW7 regulatory module determines grain shape and simultaneously improves rice yield and grain quality. Nat Genet, 47(8): 949-954. |
| [29] | Wang Y X, Xiong G S, Hu J, Jiang L, Yu H, Xu J, Fang Y X, Zeng L J, Xu E B, Xu J, Ye W J, Meng X B, Liu R F, Chen H Q, Jing Y H, Wang Y H, Zhu X D, Li J Y, Qian Q.2015. Copy number variation at theGL7 locus contributes to grain size diversity in rice. Nat Genet, 47(8): 944-948. |
| [30] | Wu W G, Liu X Y, Wang M H, Meyer R S, Luo X J, Ndjiondjop M N, Tan L B, Zhang J W, Wu J Z, Cai H W, Sun C Q, Wang X K, Wing R A, Zhu Z F.2017. A single-nucleotide polymorphism causes smaller grain size and loss of seed shattering during African rice domestication.Nat Plants, 3: 17064. |
| [31] | Wu W X, Zheng X M, Lu G W, Zhong Z Z, Gao H, Chen L P, Wu C Y, Wang H J, Wang Q, Zhou K N, Wang J L, Wu F Q, Zhang X, Guo X P, Cheng Z J, Lei C L, Lin Q B, Jiang L, Wang H Y, Ge S, Wan J M.2013. Association of functional nucleotide polymorphisms at DTH2 with the northward expansion of rice cultivation in Asia. Proc Natl Acad Sci USA, 110(8): 2775-2780. |
| [32] | Xia D, Zhou H, Liu R J, Dan W H, Li P B, Wu B, Chen J X, Wang L Q, Gao G J, Zhang Q L, He Y Q.2018. GL3.3, a novel QTL encoding a GSK3/SHAGGY-like kinase, epistatically interacts with GS3 to form extra-long grains in rice. Mol Plant, 11(5): 754-756. |
| [33] | Xu C J, Liu Y, Li Y B, Xu X D, Xu C G, Li X H, Xiao J H, Zhang Q F.2015. Differential expression ofGS5 regulates grain size in rice. J Exp Bot, 66(9): 2611-2623. |
| [34] | Ying J Z, Ma M, Bai C, Huang X H, Liu J L, Fang Y Y, Song X J.2018. TGW3, a major QTL that negatively modulates grain length and weight in rice. Mol Plant, 11(5): 750-753. |
| [35] | Yu J P, Xiong H Y, Zhu X Y, Zhang H L, Li H H, Miao J L, Wang W S, Tang Z S, Zhang Z Y, Yao G X, Zhang Q, Pan Y H, Wang X, Rashid M A R, Li J J, Gao Y M, Li Z K, Yang W C, Fu X D, Li Z C.2017. OsLG3 contributing to rice grain length and yield was mined by Ho-LAMap. BMC Biol, 15: 28. |
| [36] | Yu J P, Miao J L, Zhang Z Y, Xiong H Y, Zhu X Y, Sun X M, Pan Y H, Liang Y T, Zhang Q, Rashid M A R, Li J J, Zhang H L, Li Z C.2018. Alternative splicing of OsLG3b controls grain length and yield in japonica rice. Plant Biotechnol J, 16(9): 1667-1678. |
| [37] | Zhang H W, Fan Y Y, Zhu Y J, Chen J Y, Yu S B, Zhuang J Y.2016. Dissection of theqTGW1.1 region into two tightly-linked minor QTLs having stable effects for grain weight in rice. BMC Genet, 17(1): 98. |
| [38] | Zhang X J, Wang J F, Huang J, Lan H X, Wang C L, Yin C F, Wu Y Y, Tang H J, Qian Q, Li J Y, Zhang H S.2012. Rare allele ofOsPPKL1 associated with grain length causes extra-large grain and a significant yield increase in rice. Proc Natl Acad Sci USA, 109: 21534-21539. |
| [39] | Zhao D S, Li Q F, Zhang C Q, Zhang C, Yang Q Q, Pan L X, Ren X Y, Lu J, Gu M H, Liu Q Q.2018. GS9 acts as a transcriptional activator to regulate rice grain shape and appearance quality. Nat Commun, 9: 1240. |
| [40] | Zheng K L, Huang N, Bennett J, Khush G S.1995. PCR-ased marker-assisted selection in rice breeding. IRRI Discussion Paper Series No. 12. Los Banos, the Phillipines:International Rice Research Institute. |
| [41] | Zhu Y J, Fan Y Y, Wang K, Huang D R, Liu W Z, Ying J Z, Zhuang J Y.2017. Rice Flowering LocusT1 plays an important role in heading date influencing yield traits in rice. Sci Rep, 7: 4918. |
| [42] | Zuo J R, Li J Y.2014. Molecular genetic dissection of quantitative trait loci regulating rice grain size.Annu Rev Genet, 48: 99-118. |
| No related articles found! |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||