
Rice Science ›› 2015, Vol. 22 ›› Issue (5): 217-226.DOI: 10.1016/S1672-6308(14)60302-4
• Orginal Article • Previous Articles Next Articles
Ya-fang Zhang1, Yu-yin Ma2, Zong-xiang Chen1, Jie Zou1, Tian-xiao Chen1, Qian-qian Li1, Xue-biao Pan1, Shi-min Zuo1( )
)
Received:2015-04-21
Accepted:2015-07-01
Online:2015-05-15
Published:2015-07-24
Ya-fang Zhang, Yu-yin Ma, Zong-xiang Chen, Jie Zou, Tian-xiao Chen, Qian-qian Li, Xue-biao Pan, Shi-min Zuo. Genome-Wide Association Studies Reveal New Genetic Targets for Five Panicle Traits of International Rice Varieties[J]. Rice Science, 2015, 22(5): 217-226.
Add to citation manager EndNote|Ris|BibTeX
URL: http://www.ricesci.org/EN/10.1016/S1672-6308(14)60302-4
| Trait | Location | ADM (49) | ARO (8) | AUS (52) | IND (53) | TEJ (78) | TRJ (75) |
|---|---|---|---|---|---|---|---|
| Panicle length (cm) | Arkansas, America | 24.6 ± 3.6 A | 30.3 ± 2.0 A | 24.9 ± 2.8 A | 25.8 ± 2.6 A | 21.3 ± 2.7 A | 25.0 ± 3.1 A |
| Yangzhou, China | 27.9 ± 4.7 B | 32.8 ± 4.1 A | 28.2 ± 3.5 B | 27.1 ± 3.7 A | 24.5 ± 4.7 B | 30.1 ± 4.5 B | |
| Primary branch number | Arkansas, America | 9.9 ± 1.7 A | 8.4 ± 1.2 A | 8.6 ± 1.3 A | 9.9 ± 1.3 A | 9.7 ± 1.5 A | 11.2 ± 1.8 A |
| Yangzhou, China | 12.1 ± 2.2 B | 13.1 ± 0.8 B | 12.1 ± 1.9 B | 12.8 ± 1.7 B | 11.4 ± 2.4 B | 14.3 ± 2.4 B | |
| Grain length (cm) | Arkansas, America | 8.6 ± 0.9 A | 9.4 ± 0.6 A | 8.0 ± 0.8 A | 8.2 ± 0.7 A | 7.9 ± 0.8 A | 9.0 ± 0.8 A |
| Yangzhou, China | 8.7 ± 0.9 A | 9.4 ± 0.7 A | 8.2 ± 0.7 A | 8.3 ± 0.7 A | 7.9 ± 0.8 A | 9.0 ± 0.7 A | |
| Grain width (mm) | Arkansas, America | 3.2 ± 0.3 A | 2.6 ± 0.3 A | 2.9 ± 0.3 A | 2.9 ± 0.2 A | 3.5 ± 0.2 A | 3.0 ± 0.4 A |
| Yangzhou, China | 3.2 ± 0.4 A | 2.7 ± 0.3 A | 2.9 ± 0.4 A | 3.0 ± 0.3 A | 3.5 ± 0.3 A | 3.0 ± 0.4 A | |
| Grain length/width ratio | Arkansas, America | 2.7 ± 0.5 A | 3.6 ± 0.5 A | 2.8 ± 0.5 A | 2.8 ± 0.4 A | 2.2 ± 0.3 A | 3.0 ± 0.5 A |
| Yangzhou, China | 2.7 ± 0.5 A | 3.5 ± 0.5 A | 2.9 ± 0.6 A | 2.8 ± 0.5 A | 2.3 ± 0.3 A | 3.1 ± 0.5 A |
Table 1 Differences in panicle traits of different sub-group varieties in two locations revealed by t-test.
| Trait | Location | ADM (49) | ARO (8) | AUS (52) | IND (53) | TEJ (78) | TRJ (75) |
|---|---|---|---|---|---|---|---|
| Panicle length (cm) | Arkansas, America | 24.6 ± 3.6 A | 30.3 ± 2.0 A | 24.9 ± 2.8 A | 25.8 ± 2.6 A | 21.3 ± 2.7 A | 25.0 ± 3.1 A |
| Yangzhou, China | 27.9 ± 4.7 B | 32.8 ± 4.1 A | 28.2 ± 3.5 B | 27.1 ± 3.7 A | 24.5 ± 4.7 B | 30.1 ± 4.5 B | |
| Primary branch number | Arkansas, America | 9.9 ± 1.7 A | 8.4 ± 1.2 A | 8.6 ± 1.3 A | 9.9 ± 1.3 A | 9.7 ± 1.5 A | 11.2 ± 1.8 A |
| Yangzhou, China | 12.1 ± 2.2 B | 13.1 ± 0.8 B | 12.1 ± 1.9 B | 12.8 ± 1.7 B | 11.4 ± 2.4 B | 14.3 ± 2.4 B | |
| Grain length (cm) | Arkansas, America | 8.6 ± 0.9 A | 9.4 ± 0.6 A | 8.0 ± 0.8 A | 8.2 ± 0.7 A | 7.9 ± 0.8 A | 9.0 ± 0.8 A |
| Yangzhou, China | 8.7 ± 0.9 A | 9.4 ± 0.7 A | 8.2 ± 0.7 A | 8.3 ± 0.7 A | 7.9 ± 0.8 A | 9.0 ± 0.7 A | |
| Grain width (mm) | Arkansas, America | 3.2 ± 0.3 A | 2.6 ± 0.3 A | 2.9 ± 0.3 A | 2.9 ± 0.2 A | 3.5 ± 0.2 A | 3.0 ± 0.4 A |
| Yangzhou, China | 3.2 ± 0.4 A | 2.7 ± 0.3 A | 2.9 ± 0.4 A | 3.0 ± 0.3 A | 3.5 ± 0.3 A | 3.0 ± 0.4 A | |
| Grain length/width ratio | Arkansas, America | 2.7 ± 0.5 A | 3.6 ± 0.5 A | 2.8 ± 0.5 A | 2.8 ± 0.4 A | 2.2 ± 0.3 A | 3.0 ± 0.5 A |
| Yangzhou, China | 2.7 ± 0.5 A | 3.5 ± 0.5 A | 2.9 ± 0.6 A | 2.8 ± 0.5 A | 2.3 ± 0.3 A | 3.1 ± 0.5 A |
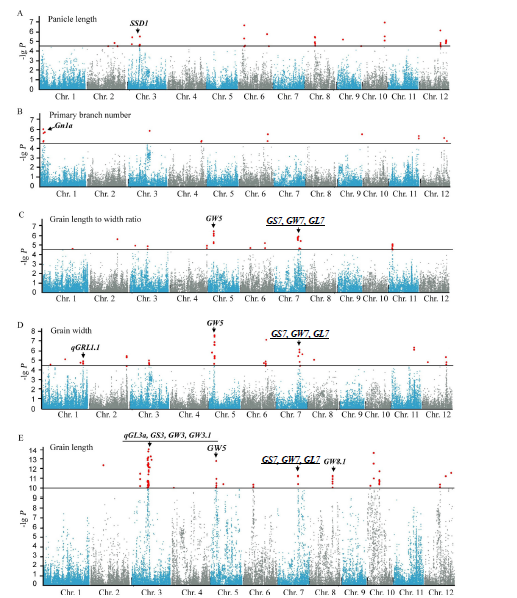
Fig. 2. Genome-wide association studies of 5 panicle traits using 315 rice varieties. (A to E, Manhattan plots of mixed linear model for five traits. Black horizontal lines indicate the genome-wide significance threshold.)
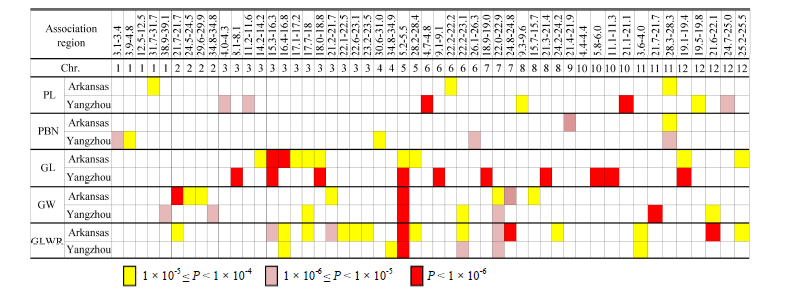
Fig. 3. Comparison of associated regions of five panicle traits detected in two locations. (PL, Panicle length; PBN, Primary branch number; GL, Grain length; GW, Grain width; GLWR, Grain length to width ratio.)
| Trait | SNP ID | Chromosome | Position (bp) | Major allele | Minor allele | Minor allele frequency | P value | Marker R2 | Phenotypic difference |
|---|---|---|---|---|---|---|---|---|---|
| between alleles | |||||||||
| PL | id3002504 | 3 | 4 364 855 | A | G | 0.31 | 3.24E-06 | 7.38 | 4.23 |
| id3006008 | 3 | 11 627 714 | T | C | 0.22 | 2.70E-06 | 8.88 | 4.94 | |
| id6003381 | 6 | 4 846 082 | A | T | 0.37 | 1.90E-07 | 8.04 | 4.3 | |
| id8003039 | 8 | 9 380 990 | C | T | 0.26 | 3.02E-06 | 6.5 | 4.46 | |
| ud10001221 | 10 | 21 175 430 | G | C | 0.14 | 9.57E-08 | 8.35 | 5.57 | |
| id12006578 | 12 | 19 520 927 | T | C | 0.5 | 6.53E-07 | 7.32 | 1.74 | |
| id12009010 | 12 | 24 997 487 | A | C | 0.31 | 6.84E-06 | 6.04 | 4.35 | |
| PBN | id1002477 | 1 | 3 096 543 | A | T | 0.26 | 9.76E-07 | 7.14 | -2.27 |
| id1003789 | 1 | 4 545 468 | C | T | 0.28 | 2.01E-06 | 6.66 | -2.15 | |
| id4010535 | 4 | 31 070 028 | A | G | 0.37 | 1.57E-05 | 5.76 | -1.9 | |
| id6014585 | 6 | 26 364 670 | A | T | 0.29 | 3.26E-06 | 6.44 | 1.99 | |
| id11011548 | 11 | 28 322 308 | T | G | 0.07 | 4.94E-06 | 6.23 | 3.02 | |
| GL | ud3000463 | 3 | 8113 668 | A | T | 0.45 | 3.46E-12 | 8.74 | 0.32 |
| id3008053 | 3 | 16 074 273 | C | T | 0.36 | 9.41E-15 | 11.79 | 0.64 | |
| id3009175 | 3 | 18 788 035 | G | A | 0.27 | 1.27E-13 | 11.25 | 0.9 | |
| id5002706 | 5 | 5 339 063 | C | T | 0.24 | 1.69E-13 | 10.29 | 0.93 | |
| wd6000477 | 6 | 9 123 484 | C | T | 0.18 | 4.78E-11 | 7.59 | 1.01 | |
| ud7001337 | 7 | 18 964 116 | G | C | 0.28 | 5.56E-12 | 9.05 | 0.87 | |
| id8005966 | 8 | 21 416 851 | G | T | 0.27 | 7.11E-12 | 8.88 | 0.83 | |
| id10001788 | 10 | 5 878 902 | A | G | 0.26 | 3.18E-13 | 11.34 | 0.95 | |
| wd10002564 | 10 | 11 284 071 | C | T | 0.16 | 1.98E-12 | 9.07 | 1.09 | |
| id12006498 | 12 | 19 402 914 | C | A | 0.43 | 6.25E-12 | 9.33 | -0.84 | |
| GW | id1024648 | 1 | 38 923 066 | A | T | 0.41 | 1.55E-05 | 6.35 | 0.47 |
| id2015723 | 2 | 34 819 747 | T | C | 0.28 | 2.88E-06 | 6.47 | 0.33 | |
| id3008699 | 3 | 17 806 411 | G | C | 0.49 | 8.27E-06 | 6.01 | -0.48 | |
| id5002708 | 5 | 5 340 574 | A | T | 0.11 | 1.93E-08 | 9.35 | 0.48 | |
| id6011831 | 6 | 22 951 051 | C | T | 0.13 | 5.71E-08 | 11.95 | 0.55 | |
| id7003644 | 7 | 22 056 571 | A | G | 0.44 | 6.02E-07 | 7.34 | 0.46 | |
| dd11000032 | 11 | 21 706 131 | A | G | 0.19 | 3.68E-07 | 7.63 | -0.18 | |
| id12007136 | 12 | 21 630 274 | G | A | 0.06 | 3.63E-06 | 6.43 | 0.56 | |
| GLWR | wd3000595 | 3 | 16 759 297 | T | C | 0.23 | 1.30E-05 | 6.01 | 0.1 |
| id4012361 | 4 | 34 949 006 | G | A | 0.15 | 1.10E-05 | 5.71 | -0.03 | |
| id5002751 | 5 | 5 396 733 | G | C | 0.47 | 3.42E-07 | 7.65 | 0.65 | |
| id6011831 | 6 | 22 951 051 | C | T | 0.13 | 6.04E-06 | 8.47 | -0.83 | |
| id7003917 | 7 | 22 787 412 | A | G | 0.12 | 1.24E-06 | 6.95 | -0.77 | |
| id11001567 | 11 | 3 956 750 | A | G | 0.45 | 7.99E-06 | 6.09 | -0.39 |
Table 2 Genome-wide significant association of rice panicle traits.
| Trait | SNP ID | Chromosome | Position (bp) | Major allele | Minor allele | Minor allele frequency | P value | Marker R2 | Phenotypic difference |
|---|---|---|---|---|---|---|---|---|---|
| between alleles | |||||||||
| PL | id3002504 | 3 | 4 364 855 | A | G | 0.31 | 3.24E-06 | 7.38 | 4.23 |
| id3006008 | 3 | 11 627 714 | T | C | 0.22 | 2.70E-06 | 8.88 | 4.94 | |
| id6003381 | 6 | 4 846 082 | A | T | 0.37 | 1.90E-07 | 8.04 | 4.3 | |
| id8003039 | 8 | 9 380 990 | C | T | 0.26 | 3.02E-06 | 6.5 | 4.46 | |
| ud10001221 | 10 | 21 175 430 | G | C | 0.14 | 9.57E-08 | 8.35 | 5.57 | |
| id12006578 | 12 | 19 520 927 | T | C | 0.5 | 6.53E-07 | 7.32 | 1.74 | |
| id12009010 | 12 | 24 997 487 | A | C | 0.31 | 6.84E-06 | 6.04 | 4.35 | |
| PBN | id1002477 | 1 | 3 096 543 | A | T | 0.26 | 9.76E-07 | 7.14 | -2.27 |
| id1003789 | 1 | 4 545 468 | C | T | 0.28 | 2.01E-06 | 6.66 | -2.15 | |
| id4010535 | 4 | 31 070 028 | A | G | 0.37 | 1.57E-05 | 5.76 | -1.9 | |
| id6014585 | 6 | 26 364 670 | A | T | 0.29 | 3.26E-06 | 6.44 | 1.99 | |
| id11011548 | 11 | 28 322 308 | T | G | 0.07 | 4.94E-06 | 6.23 | 3.02 | |
| GL | ud3000463 | 3 | 8113 668 | A | T | 0.45 | 3.46E-12 | 8.74 | 0.32 |
| id3008053 | 3 | 16 074 273 | C | T | 0.36 | 9.41E-15 | 11.79 | 0.64 | |
| id3009175 | 3 | 18 788 035 | G | A | 0.27 | 1.27E-13 | 11.25 | 0.9 | |
| id5002706 | 5 | 5 339 063 | C | T | 0.24 | 1.69E-13 | 10.29 | 0.93 | |
| wd6000477 | 6 | 9 123 484 | C | T | 0.18 | 4.78E-11 | 7.59 | 1.01 | |
| ud7001337 | 7 | 18 964 116 | G | C | 0.28 | 5.56E-12 | 9.05 | 0.87 | |
| id8005966 | 8 | 21 416 851 | G | T | 0.27 | 7.11E-12 | 8.88 | 0.83 | |
| id10001788 | 10 | 5 878 902 | A | G | 0.26 | 3.18E-13 | 11.34 | 0.95 | |
| wd10002564 | 10 | 11 284 071 | C | T | 0.16 | 1.98E-12 | 9.07 | 1.09 | |
| id12006498 | 12 | 19 402 914 | C | A | 0.43 | 6.25E-12 | 9.33 | -0.84 | |
| GW | id1024648 | 1 | 38 923 066 | A | T | 0.41 | 1.55E-05 | 6.35 | 0.47 |
| id2015723 | 2 | 34 819 747 | T | C | 0.28 | 2.88E-06 | 6.47 | 0.33 | |
| id3008699 | 3 | 17 806 411 | G | C | 0.49 | 8.27E-06 | 6.01 | -0.48 | |
| id5002708 | 5 | 5 340 574 | A | T | 0.11 | 1.93E-08 | 9.35 | 0.48 | |
| id6011831 | 6 | 22 951 051 | C | T | 0.13 | 5.71E-08 | 11.95 | 0.55 | |
| id7003644 | 7 | 22 056 571 | A | G | 0.44 | 6.02E-07 | 7.34 | 0.46 | |
| dd11000032 | 11 | 21 706 131 | A | G | 0.19 | 3.68E-07 | 7.63 | -0.18 | |
| id12007136 | 12 | 21 630 274 | G | A | 0.06 | 3.63E-06 | 6.43 | 0.56 | |
| GLWR | wd3000595 | 3 | 16 759 297 | T | C | 0.23 | 1.30E-05 | 6.01 | 0.1 |
| id4012361 | 4 | 34 949 006 | G | A | 0.15 | 1.10E-05 | 5.71 | -0.03 | |
| id5002751 | 5 | 5 396 733 | G | C | 0.47 | 3.42E-07 | 7.65 | 0.65 | |
| id6011831 | 6 | 22 951 051 | C | T | 0.13 | 6.04E-06 | 8.47 | -0.83 | |
| id7003917 | 7 | 22 787 412 | A | G | 0.12 | 1.24E-06 | 6.95 | -0.77 | |
| id11001567 | 11 | 3 956 750 | A | G | 0.45 | 7.99E-06 | 6.09 | -0.39 |
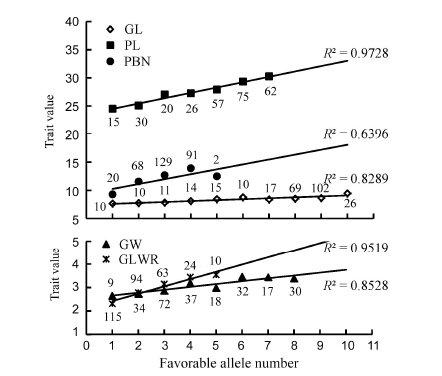
Fig. 4. Linear regression analysis on relationship between the average trait value and the number of favorable alleles.( The number located under each trait mark indicates the number of favorable alleles. R2 indicates the regression coefficient between the trait value and the number of favorable alleles controlling this trait. PL, Panicle length (cm); PBN, Primary branch number; GL, Grainlength (cm); GW, Grain width (cm); GLWR, Grain length to width ratio..)
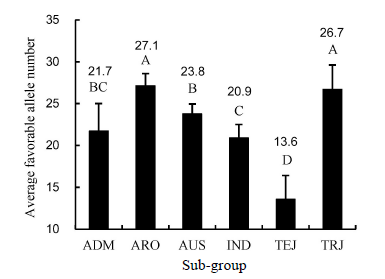
Fig. 5. Comparison of number of favorable alleles associated with five panicle traits of different sub-groups.( The number above the column indicates the number of favorable alleles in each sub-group. Different capital letters above the column indicate significant statistical difference at 0.01 level. ADM, Admix; ARO, Aromatic; AUS, Aus; IND, Indica; TEJ,Temperate japonica; TRJ, Tropical japonica.)
| [1] | Asano K, Miyao A, Hirochika H, Kitano H, Matsuoka M, Ashikari M.2010. SSD1, which encodes a plant-specific novel protein, controls plant elongation by regulating cell division in rice. Proc Jpn Acad Ser B Phys Biol Sci, 86(3): 265-273. |
| [2] | Bradbury P J, Zhang Z, Kroon D E, Casstevens T M, Ramdoss Y, Buckler E S.2007. TASSEL: Software for association mapping of complex traits in diverse samples.Bioinformatics, 23: 2633-2635. |
| [3] | Dang X J, Thi T G T, Dong G S, Wang H, Edzesi W H, Hong D.2014. Genetic diversity and association mapping of seed vigor in rice (Oryza sativa L.).Planta, 239: 1309-1319. |
| [4] | de Oliveira Borba T C, Brondani R P V, Breseghello F, Coelho A S G, Mendonca J A, Rangel P H N, Brondani C.2010. Association mapping for yield and grain quality traits in rice (Oryza sativa L.). Genet Mol Biol, 33: 515-524. |
| [5] | Han B, Huang X H.2013. Sequencing-based genome-wide association study in rice.Curr Opin Plant Biol, 16: 133-138. |
| [6] | Huang R Y, Jiang L R, Zheng J S, Wang T S, Wang H C, Huang Y M, Hong Z L.2013. Genetic bases of rice grain shape: So many genes, so little known.Trends Plant Sci, 18(4): 218-226. |
| [7] | Huang X H, Wei X H, Sang T, Zhao Q, Feng Q, Zhao Y, Li C Y, Zhu C R, Lu T T, Zhang Z W, Li M, Fan D L, Guo Y L, Wang A H, Wang L, Deng L W, Li W J, Lu Y Q, Weng Q J, Liu K Y, Huang T, Zhou T Y, Jing Y F, Li W, Lin Z, Buckler E S, Qian Q, Zhang Q F, Li J Y, Han B.2010. Genome-wide association studies of 14 agronomic traits in rice landraces.Nat Genet, 42: 961-967. |
| [8] | Ikeda M, Miura K, Aya K, Kitano H, Matsuoka M.2013. Genes offering the potential for designing yield-related traits in rice. Curr Opin Plant Biol, 16: 213-220. |
| [9] | Jin L, Lu Y, Xiao P, Sun M, Corke H, Bao J S.2010. Genetic diversity and population structure of a diverse set of rice germplasm for association mapping.Theor Appl Genet, 121: 475-487. |
| [10] | Liu G F, Yang J, Zhu J.2006. Mapping QTL for biomass yield and its components in rice (Oryza sativa L.).Acta Genet Sin, 33(7): 607-616. |
| [11] | Mao H L, Sun S Y, Yao J, Wang C R, Yu S B, Xu C G, Li X H, Zhang Q F.2010. Linking differential domain functions of the GS3 protein to natural variation of grain size in rice.Proc Natl Acad Sci USA, 107: 19579-19584. |
| [12] | Pan X B, Liang G H, Chen Z X, Zhang Y F.2005. Breeding strategy on resistance to rice stripe disease in Jiangsu Province.Jiangsu Agric Sci, 5: 22-23. (in Chinese with English abstract) |
| [13] | Singh R, Singh A K, Sharma T R, Singh A, Singh N K.2012. Fine mapping of grain length QTLs on chromosomes 1 and 7 in Basmati rice (Oryza sativa L.).J Plant Biochem Biotechnol, 21: 157-166. |
| [14] | Shao G N, Wei X J, Chen M L, Tang S Q, Luo J, Jiao G A, Xie L H, Hu P S.2012. Allelic variation for a candidate gene for GS7, responsible for grain shape in rice.Theor Appl Genet, 125: 1303-1312. |
| [15] | Shomura A, Izawa T, Ebana K, Ebitani T, Kanegae H, Konishi S, Yano M.2008. Deletion in a gene associated with grain size increased yields during rice domestication.Nat Genet, 40: 1023-1028. |
| [16] | Sreedhar S, Reddy T D, Ramesha M S.2011. Genotype × environment interaction and stability for yield and its components in hybrid rice cultivars (Oryza sativa L.). Int J Plant Breeding Genet, 5: 194-208. |
| [17] | Tran T T G, Dang X J, Liu Q M, Zhao K M, Wang H, Hong D L.2014. Association analysis of rice grain traits with SSR markers.Chin J Rice Sci, 28(3): 243-257. (in Chinese with English abstract) |
| [18] | Wang C L.2005. The current situation and development trend of rice breeding in Jiangsu Province.Jiangsu Agric Sci, 33(2): 1-6. (in Chinese with English abstract) |
| [19] | Wang S K, Li S, Liu Q, Wu K, Zhang J Q, Wang S S, Wang Y, Chen X B, Zhang Y, Gao C X, Wang F, Huang H X, Fu X D.2015. The OsSPL16-GW7 regulatory module determines grain shape and simultaneously improves rice yield and grain quality.Nat Genet, 47(8): 949-954. |
| [20] | Wang Y X, Xiong G S, Hu J, Jiang L, Yu H, Xu J, Fang Y X, Zeng L J, Xu E B, Xu J, Ye W J, Meng X B, Liu R F, Chen H Q, Jing Y H, Wang Y H, Zhu X D, Li J Y, Qian Q.2015. Copy number variation at the GL7 locus contributes to grain size diversity in rice.Nat Genet, 47(8): 944-948. |
| [21] | Wei X H, Yuan X P, Yu H Y, Wang Y P, Xu Q, Tang S X.2009. SSR analysis of genetic variation in Chinese major indred rice varieties.Chin J Rice Sci, 23(3): 237-244. (in Chinese with English abstract) |
| [22] | Wei X H, Tang S X, Yu H Y, Wang Y P, Yuan X P, Xu Q.2010. Beneficial analysis on introduced rice germplasm from abroad in China.Chin J Rice Sci, 24(1): 5-11. (in Chinese with English abstract) |
| [23] | Xie X B, Song M H, Jin F X, Ahn S N, Suh J P, Hwang H G, McCouch S R.2006. Fine mapping of a grain weight quantitative trait locus on rice chromosome 8 using near-isogenic lines derived from a cross between Oryza sativa and Oryza rufipogon. Theor Appl Genet, 113: 885-894. |
| [24] | Xuan Y S, Jiang W Z, Liu X H, Cheng Z H, Koh H J, Yuan D L.2010. Comparative analysis of genetic diversity of commercial rice cultivars in northeastern China.J Plant Genet Res, 11(2): 206-212. (in Chinese with English abstract) |
| [25] | Zhang H G, Zhu G Y, Feng Z Q, Xu M, Ji J A, Pei Y, Qian K, Tang S Z, Gu M H.2014. Analysis on yield and quality of the late-maturity medium japonica rice varieties released in Jiangsu Province in the last 30 years.Chin J Rice Sci, 28(3): 327-334. (in Chinese with English abstract) |
| [26] | Zhang Q, Yao G X, Hu G L, Chen C, Tang B, Zhang H L, Li Z C.2012. Fine mapping of qTGW3-1, a QTL for 1000-grain weight on chromosome 3 in rice.J Intl Agric, 11(6): 879-887. |
| [27] | Zhao K Y, Tung C W, Eizenga G C, Wright M H, Ali M L, Price A H, Jnorton G J, Islam M R, Reynolds A, Mezey J, Mcclung A M, Bustamante C D, McCouch S R.2011. Genome-wide association mapping reveals a rich genetic architecture of complex traits in Oryza sativa.Nat Commun, DOI: 10.1038/ncomms1467. |
| [28] | Zheng W J, Liu X.2009. Research progress and prospects of rice stripe virus (RSV).China Plant Prot, 5: 12-15. (in Chinese with English abstract) |
| [29] | Zhou S C, Wang J S, Li H, Huang D Q, Xie Z W.2001. Review and reflection of rice breeding in China.China Rice, 7(2): 5-7. (in Chinese) |
| [30] | Zhou Y Y, Sha A Q, Fan B G, Wang J, Gong J L, Long H Y, Hu Y J, Ming Q L, Liu G L, Luo X C, Zhu F H.2012. Studies on super-high yield formation rule and cultivation approach of indica-japonica hybrid rice ‘Yongyou 8’.Jiangsu Agric Sci, 40(2): 45-47. (in Chinese with English abstract) |
| [31] | Zhu L H.2007. Some critical considerations on rice high-yielding breeding in China. J Nanjing Agric Univ, 30(1): 129-135. (in Chinese with English abstract) |
| [1] | Amrit Kumar Nayak, Anilkumar C, Sasmita Behera, Rameswar Prasad Sah, Gera Roopa Lavanya, Awadhesh Kumar, Lambodar Behera, Muhammed Azharudheen Tp. Genetic Dissection of Grain Size Traits Through Genome-Wide Association Study Based on Genic Markers in Rice [J]. Rice Science, 2022, 29(5): 462-472. |
| [2] | Rakotoson Tatiana, Dusserre Julie, Letourmy Philippe, Frouin Julien, Ramonta Ratsimiala Isabelle, Victorine Rakotoarisoa Noronirina, Cao Tuong-Vi, Vom Brocke Kirsten, Ramanantsoanirina Alain, Ahmadi Nourollah, Raboin Louis-Marie. Genome-Wide Association Study of Nitrogen Use Efficiency and Agronomic Traits in Upland Rice [J]. Rice Science, 2021, 28(4): 379-390. |
| [3] | Yang Lv, Yueying Wang, Jahan Noushin, Haitao Hu, Ping Chen, Lianguang Shang, Haiyan Lin, Guojun Dong, Jiang Hu, Zhenyu Gao, Qian Qian, Yu Zhang, Longbiao Guo. Genome-Wide Association Analysis and Allelic Mining of Grain Shape-Related Traits in Rice [J]. Rice Science, 2019, 26(6): 384-392. |
| [4] | Zongxiang Chen, Zhiming Feng, Houxiang Kang, Jianhua Zhao, Tianxiao Chen, Qianqian Li, Hongbing Gong, Yafang Zhang, Xijun Chen, Xuebiao Pan, Wende Liu, Guoliang Wang, Shimin Zuo. Identification of New Resistance Loci Against Sheath Blight Disease in Rice Through Genome-Wide Association Study [J]. Rice Science, 2019, 26(1): 21-31. |
| [5] | Lee Jae-Sung, Wissuwa Matthias, B. Zamora Oscar, M. Ismail Abdelbagi. Novel Sources of aus Rice for Zinc Deficiency Tolerance Identified Through Association Analysis Using High-Density SNP Array [J]. Rice Science, 2018, 25(5): 293-296. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||