
Rice Science ›› 2015, Vol. 22 ›› Issue (4): 162-170.DOI: 10.1016/S1672-6308(14)60299-7
• Orginal Article • Previous Articles Next Articles
Zhong-hua Sheng, Peng-fei Fang, San-feng Li, Gui-ai Jiao, Li-hong Xie, Pei-song Hu, Shao-qing Tang, Xiang-jin Wei( )
)
Received:2015-04-02
Accepted:2015-05-04
Online:2015-07-28
Published:2015-05-27
About author:Corresponding author:WEI Xiang-jin(weixj99@126.com)
Zhong-hua Sheng, Peng-fei Fang, San-feng Li, Gui-ai Jiao, Li-hong Xie, Pei-song Hu, Shao-qing Tang, Xiang-jin Wei. Phenotype of Rice Floury Endosperm Mutant flo7 and Fine Mapping of Mutated Gene[J]. Rice Science, 2015, 22(4): 162-170.
Add to citation manager EndNote|Ris|BibTeX
URL: http://www.ricesci.org/EN/10.1016/S1672-6308(14)60299-7
| Primer | Forward (5'-3') | Reverse (5'-3') | Product size (bp) |
| RM6411 | GGGTATTGTCGGTGTTCAGG | GAAGGCTGTACCATCCAACG | 140 |
| RM1227 | CATGGTAGCACACACCCTTG | CATCGACATGTGGACCACTC | 137 |
| RM28746 | GAAGAAAGAAGACGCCAAGAAACG | CATTCCATTCCCTTCCTCTTCG | 158 |
| C5-2 | GCTGTCAGATCGGTAGGCA | ACCACATCAACCCAGAAAA | 90 |
| C5-15 | TGTTTGTGCCTTCAGATAA | TGAGATAGGGGAAGTAGAT | 87 |
| C2-6 | AGATTCTTTTCCAAACAAC | AATCATCATATTGCCGAAA | 81 |
| C2-11 | TGCATCATCCAATCTGTCTTGG | TCGACAAGCCAATCTACTCTTCC | 139 |
| C7-3 | CTACGTGCATCCATCGAGAA | TGAATTGTTTACCGGAACCC | 153 |
| C7-5 | TCTTCTACTCCTCGTCTTCGTTCG | CATGCAGAGCAGAGACTTCTTGG | 143 |
| C7-12 | TGCTGGACACAGCAAGTTTT | TAGATGGTGGGGGTGATAGG | 185 |
| C7-15 | GCGTTGATTGATGATGATGG | CGACCCGACGTAAAAGAGAA | 113 |
Table 1 Primers for fine mapping of Flo7.
| Primer | Forward (5'-3') | Reverse (5'-3') | Product size (bp) |
| RM6411 | GGGTATTGTCGGTGTTCAGG | GAAGGCTGTACCATCCAACG | 140 |
| RM1227 | CATGGTAGCACACACCCTTG | CATCGACATGTGGACCACTC | 137 |
| RM28746 | GAAGAAAGAAGACGCCAAGAAACG | CATTCCATTCCCTTCCTCTTCG | 158 |
| C5-2 | GCTGTCAGATCGGTAGGCA | ACCACATCAACCCAGAAAA | 90 |
| C5-15 | TGTTTGTGCCTTCAGATAA | TGAGATAGGGGAAGTAGAT | 87 |
| C2-6 | AGATTCTTTTCCAAACAAC | AATCATCATATTGCCGAAA | 81 |
| C2-11 | TGCATCATCCAATCTGTCTTGG | TCGACAAGCCAATCTACTCTTCC | 139 |
| C7-3 | CTACGTGCATCCATCGAGAA | TGAATTGTTTACCGGAACCC | 153 |
| C7-5 | TCTTCTACTCCTCGTCTTCGTTCG | CATGCAGAGCAGAGACTTCTTGG | 143 |
| C7-12 | TGCTGGACACAGCAAGTTTT | TAGATGGTGGGGGTGATAGG | 185 |
| C7-15 | GCGTTGATTGATGATGATGG | CGACCCGACGTAAAAGAGAA | 113 |
| Primer | Accession number | Forward (5'-3') | Reverse (5'-3') |
| Actin | X16280 | CATGCTATCCCTCGTCTCGACCT | CGCACTTCATGATGGAGTTGTAT |
| OsAGPS1 | AK073146 | AGAATGCTCGTATTGGAGAAAATG | GGCAGCATGGAATAAACCAC |
| OsAGPS2a | AK071826 | ACTCCAAGAGCTCGCAGACC | GCCTGTAGTTGGCACCCAGA |
| OsAGPS2b | AK103906 | AACAATCGAAGCGCGAGAAA | GCCTGTAGTTGGCACCCAGA |
| OsAGPL1 | D50317 | GGAAGACGGATGATCGAGAAAG | CACATGAGATGCACCAACGA |
| OsAGPL3 | AK069296 | AAGCCAGCCATGACCATTTG | CACACGGTAGATTCACGAGACAA |
| OsAGPL4 | AK121036 | TCAACGTCGATGCAGCAAAT | ATCCCTCAGTTCCTAGCCTCATT |
| OsSSI | D16202 | GGGCCTTCATGGATCAACC | CCGCTTCAAGCATCCTCATC |
| OsSSIIa | AF419099 | GCTTCCGGTTTGTGTGTTCA | CTTAATACTCCCTCAACTCCACCAT |
| OsSSIIIa | AY100469 | GCCTGCCCTGGACTACATTG | GCAAACATATGTACACGGTTCTGG |
| OsBEIIb | D16201 | ATGCTAGAGTTTGACCGC | AGTGTGATGGATCCTGCC |
| GBSSI | AK070431 | TCCGAGAGGTTCAGGTCATC | ATGAGCTCCTCGGCGTAGTA |
| OsISA1 | AB093426 | TGCTCAGCTACTCCTCCATCATC | AGGACCGCACAACTTCAACATA |
| OsISA2 | AC132483 | TAGAGGTCCTCTTGGAGG | AATCAGCTTCTGAGTCACCG |
| OsISA3 | AP005574 | ACAGCTTGAGACACTGGGTTGAG | GCATCAAGAGGACAACCATCTG |
| OsPUL | AB012915 | ACCTTTCTTCCATGCTGG | CAAAGGTCTGAAAGATGGG |
| OsPHOL | AK063766 | TTGGCAGGAAGGTTTCGCT | CGAAGCCTGAAGTGAACTTGCT |
Table 2 Primers used to assay starch synthesis-related gene transcription.
| Primer | Accession number | Forward (5'-3') | Reverse (5'-3') |
| Actin | X16280 | CATGCTATCCCTCGTCTCGACCT | CGCACTTCATGATGGAGTTGTAT |
| OsAGPS1 | AK073146 | AGAATGCTCGTATTGGAGAAAATG | GGCAGCATGGAATAAACCAC |
| OsAGPS2a | AK071826 | ACTCCAAGAGCTCGCAGACC | GCCTGTAGTTGGCACCCAGA |
| OsAGPS2b | AK103906 | AACAATCGAAGCGCGAGAAA | GCCTGTAGTTGGCACCCAGA |
| OsAGPL1 | D50317 | GGAAGACGGATGATCGAGAAAG | CACATGAGATGCACCAACGA |
| OsAGPL3 | AK069296 | AAGCCAGCCATGACCATTTG | CACACGGTAGATTCACGAGACAA |
| OsAGPL4 | AK121036 | TCAACGTCGATGCAGCAAAT | ATCCCTCAGTTCCTAGCCTCATT |
| OsSSI | D16202 | GGGCCTTCATGGATCAACC | CCGCTTCAAGCATCCTCATC |
| OsSSIIa | AF419099 | GCTTCCGGTTTGTGTGTTCA | CTTAATACTCCCTCAACTCCACCAT |
| OsSSIIIa | AY100469 | GCCTGCCCTGGACTACATTG | GCAAACATATGTACACGGTTCTGG |
| OsBEIIb | D16201 | ATGCTAGAGTTTGACCGC | AGTGTGATGGATCCTGCC |
| GBSSI | AK070431 | TCCGAGAGGTTCAGGTCATC | ATGAGCTCCTCGGCGTAGTA |
| OsISA1 | AB093426 | TGCTCAGCTACTCCTCCATCATC | AGGACCGCACAACTTCAACATA |
| OsISA2 | AC132483 | TAGAGGTCCTCTTGGAGG | AATCAGCTTCTGAGTCACCG |
| OsISA3 | AP005574 | ACAGCTTGAGACACTGGGTTGAG | GCATCAAGAGGACAACCATCTG |
| OsPUL | AB012915 | ACCTTTCTTCCATGCTGG | CAAAGGTCTGAAAGATGGG |
| OsPHOL | AK063766 | TTGGCAGGAAGGTTTCGCT | CGAAGCCTGAAGTGAACTTGCT |
| Material | Plant height (cm) | Main panicle length (cm) | Effective tiller number per plant | No. of grains per panicle | Seed-setting rate (%) |
| WT | 67.25 ± 4.25 | 16.88 ± 2.32 | 26.25 ± 2.75 | 60.00 ± 5.80 | 66.25 ± 8.32 |
| flo7 | 74.20 ± 3.50 | 17.22 ± 1.80 | 26.75 ± 4.68 | 61.75 ± 5.25 | 82.20 ± 9.33 |
| P value (t test) | 0.003 | 0.190 | 0.150 | 0.850 | 0.036 |
Table 3 Agronomic performance of flo7 and its wild type (WT).
| Material | Plant height (cm) | Main panicle length (cm) | Effective tiller number per plant | No. of grains per panicle | Seed-setting rate (%) |
| WT | 67.25 ± 4.25 | 16.88 ± 2.32 | 26.25 ± 2.75 | 60.00 ± 5.80 | 66.25 ± 8.32 |
| flo7 | 74.20 ± 3.50 | 17.22 ± 1.80 | 26.75 ± 4.68 | 61.75 ± 5.25 | 82.20 ± 9.33 |
| P value (t test) | 0.003 | 0.190 | 0.150 | 0.850 | 0.036 |
| Material | Protein content (%) | Amylose content (%) | Gel consistency (mm) | Alkali spreading value | Hardness (N) |
| WT | 10.2 ± 0.2 | 17.9 ± 0.2 | 80.5 ± 0.5 | 6.0 ± 0.5 | 19.5 |
| flo7 | 10.7 ± 0.2 | 15.4 ± 0.3 | 56.6 ± 0.5 | 6.8 ± 0.2 | 15.4 |
| P value (t test) | 0.180 | 0.012 | 0.002 | 0.035 | 0.018 |
Table 4 Grain quality properties of flo7 and its wild type (WT).
| Material | Protein content (%) | Amylose content (%) | Gel consistency (mm) | Alkali spreading value | Hardness (N) |
| WT | 10.2 ± 0.2 | 17.9 ± 0.2 | 80.5 ± 0.5 | 6.0 ± 0.5 | 19.5 |
| flo7 | 10.7 ± 0.2 | 15.4 ± 0.3 | 56.6 ± 0.5 | 6.8 ± 0.2 | 15.4 |
| P value (t test) | 0.180 | 0.012 | 0.002 | 0.035 | 0.018 |
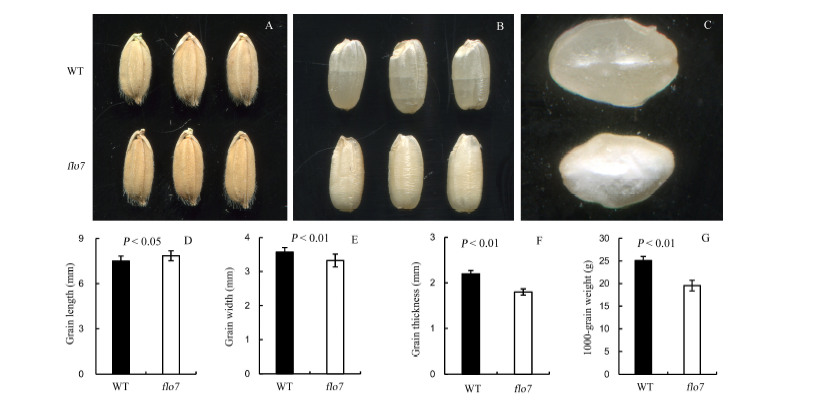
Fig. 1. Phenotypes of flo7 and its wild type (WT). A, Non-milled grains; B, Milled grains; C, Cross sections of endosperm; D, Grain length; E, Grain width; F, Grain thickness; G, 1000-grain weight. All values are given as mean ± SD.
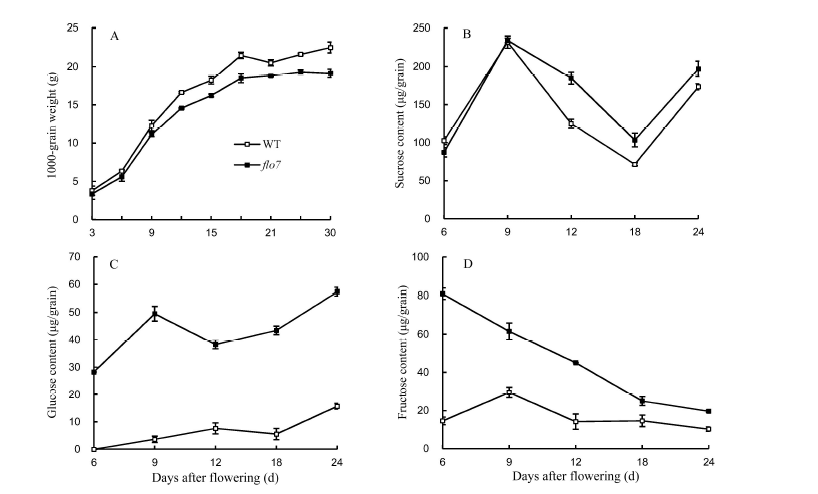
Fig. 2. Free sugar content and 1000-grain weight of flo7 and wild type (WT) during grain filling (Mean ± SD, n = 3). A, 1000-grain weight; B, Sucrose content; C, Glucose content; D, Fructose content.
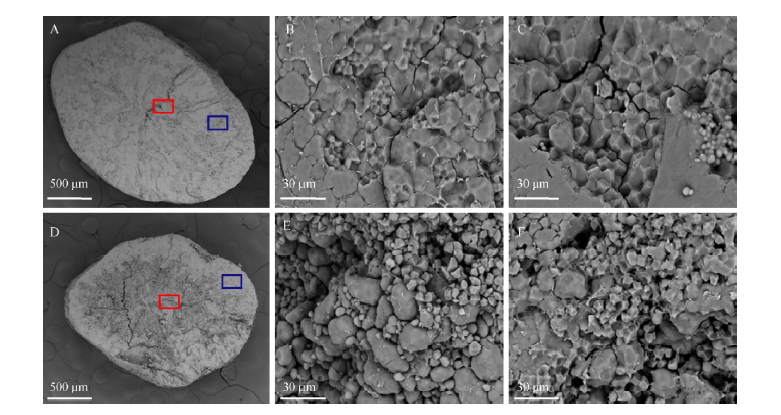

Fig. 3. Structure of starch granules in endosperm of wild type (WT) (A, B and C) and flo7 (D, E and F). A and D, Cross-sections of endosperm; B and E, Central region of mature endosperm from the cross-sections in A and D indicated by the red square; C and F, Outer portion of mature endosperm from the cross-sections in A and D indicated by the blue square.
| Combination | No. of wild type plants | No. of flo7 type plants | χ2(3:1 ) | P value |
| flo7/Nanjing 11 | 502 | 179 | 0.600 | 0.44 |
| flo7/Pei’ai 64 | 89 | 28 | 0.025 | 0.79 |
| flo7/93-11 | 102 | 36 | 0.038 | 0.77 |
Table 5 Genetic analysis of flo7.
| Combination | No. of wild type plants | No. of flo7 type plants | χ2(3:1 ) | P value |
| flo7/Nanjing 11 | 502 | 179 | 0.600 | 0.44 |
| flo7/Pei’ai 64 | 89 | 28 | 0.025 | 0.79 |
| flo7/93-11 | 102 | 36 | 0.038 | 0.77 |
| ORF | Candidate gene | Function description |
| ORF1 | Os12g0630750 | Full insert sequence |
| ORF2 | Os12g0630800 | Hypothetical conserved gene |
| ORF3 | Os12g0631025 | Non-protein coding transcript |
| ORF4 | Os12g0631100 | Small GTPase, storage protein trafficking |
| ORF5 | Os12g0631150 | Non-protein coding transcript |
| ORF6 | Os12g0631200 | Similar to ubiquitin-protein ligase / zinc ion binding protein |
| ORF7 | Os12g0631600 | Conserved hypothetical protein. |
| ORF8 | Os12g0631800 | Phytoene dehydrogenase-like |
| ORF9 | Os12g0632000 | Similar to glycine-rich RNA-binding protein 1 |
| ORF10 | Os12g0632100 | Similar to ARL2 G-protein |
| ORF11 | Os12g0632401 | Hypothetical conserved gene |
| ORF12 | Os12g0632600 | Similar to helix-loop-helix DNA-binding domain containing protein |
| ORF13 | Os12g0632700 | Malate dehydrogenase, glyoxysomal precursor |
Table 6 Candidate genes in Flo7 fine mapping interval.
| ORF | Candidate gene | Function description |
| ORF1 | Os12g0630750 | Full insert sequence |
| ORF2 | Os12g0630800 | Hypothetical conserved gene |
| ORF3 | Os12g0631025 | Non-protein coding transcript |
| ORF4 | Os12g0631100 | Small GTPase, storage protein trafficking |
| ORF5 | Os12g0631150 | Non-protein coding transcript |
| ORF6 | Os12g0631200 | Similar to ubiquitin-protein ligase / zinc ion binding protein |
| ORF7 | Os12g0631600 | Conserved hypothetical protein. |
| ORF8 | Os12g0631800 | Phytoene dehydrogenase-like |
| ORF9 | Os12g0632000 | Similar to glycine-rich RNA-binding protein 1 |
| ORF10 | Os12g0632100 | Similar to ARL2 G-protein |
| ORF11 | Os12g0632401 | Hypothetical conserved gene |
| ORF12 | Os12g0632600 | Similar to helix-loop-helix DNA-binding domain containing protein |
| ORF13 | Os12g0632700 | Malate dehydrogenase, glyoxysomal precursor |
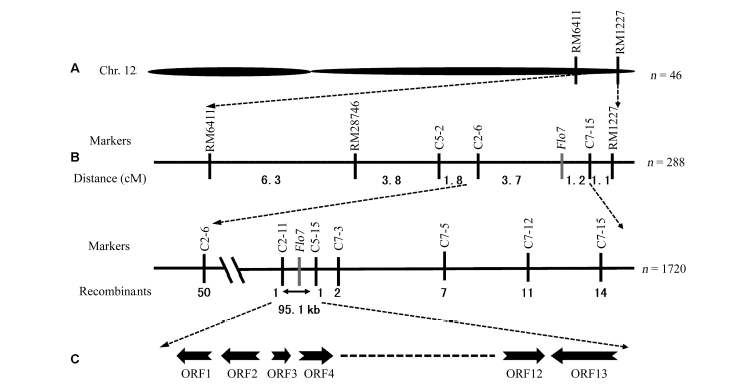
Fig. 4. Genetic mapping of gene Flo7. A, A coarse map placed the locus on chromosome 12 between RM6411 and RM1227; B, Flo7 was located in 95.1 kb region defined by the markers C2-11 and C5-15; C, Candidate region contain 13 open reading frames (ORFs).

Fig. 5. Transcription profiling of genes related to starch synthesis during grain filling (Mean ± SD, n = 3). OsAGPS1, ADP glucose pyrophosphorylase small subunit 1; OsAGPS2a, ADP glucose pyrophosphorylase small subunit 2a; OsAGPS2b, ADP glucose pyrophosphorylase small subunit 2b; OsAGPL1, ADP glucose pyrophosphorylase large subunit 1; OsAGPL3, ADP glucose pyrophosphorylase large subunit 3; OsAGPL4, ADP glucose pyrophosphorylase large subunit 4; OsSSI, Starch synthase I; OsSSIIa, Starch synthase IIa; OsSSIIIa, Starch synthase IIIa; OsGBSSI, Granule-bound starch synthase I; OsISA1, Isoamylase I; OsISA2, Isoamylase II; OsISA3, Isoamylase III; OsBEIIb, Starch branching enzyme Iib; OsPUL, Pullulanase; OsPHOL, Starch phosphorylase.
| 1 | Agricultural Industry Standard of the People’s Republic of China. 2002. The Rice Quality Measurement Method (NY147-88). Beijing, China: China Standards Press. (in Chinese) |
| 2 | Chavrier P, Goud B.1999. The role of ARF and Rab GTPases in membrane transport.Curr Opin Cell Biol, 11(4): 466-475. |
| 3 | Fu F F, Xue H W.2010. Coexpression analysis identifies Rice Starch Regulator 1, a rice AP2/EREBP family transcription factor, as a novel rice starch biosynthesis regulator.Plant Physiol, 154(2): 927-938. |
| 4 | Fukuda M, Wen L Y, Satoh-Cruz M, Kawagoe Y, Nagamura Y, Okita T W, Washida H, Sugino A, Ishino S, Ishino Y, Ogawa M, Sunada M, Ueda T, Kumamaru T.2013. A guanine nucleotide exchange factor for Rab5 proteins is essential for intracellular transport of the proglutelin from the Golgi apparatus to the protein storage vacuole in rice endosperm.Plant Physiol, 162(2): 663-674. |
| 5 | Fujita N, Yoshida M, Kondo T, Saito K, Utsumi Y, Tokunaga T, Nishi A, Satoh H, Park J H, Jane J L, Miyao A, Hirochika H, Nakamura Y.2007. Characterization of SSIIIa-deficient mutants of rice: The function of SSIIIa and pleiotropic effects by SSIIIa deficiency in the rice endosperm.Plant Physiol, 144(4): 2009-2023. |
| 6 | Gao M, Fisher D K, Kim K N, Shannon J C, Guiltinan M J.1996. Evolutionary conservation and expression patterns of maize starch branching enzyme I and IIb genes suggests isoform specialization.Plant Mol Biol, 30(6): 1223-1232. |
| 7 | Hamada S, Nozaki K, Ito H, Yoshimoto Y, Yoshida H, Hiraga S, Onodera S, Honma M, Takeda Y, Matsui H.2001. Two starch-branching-enzyme isoforms occur in different fractions of developing seeds of kidney bean.Biochem J, 359: 23-34. |
| 8 | Hanashiro I, Itoh K, Kuratomi Y, Yamazaki M, Igarashi T, Matsugasako J I, Takeda Y.2008. Granule-bound starch synthase I is responsible for biosynthesis of extra-long unit chains of amylopectin in rice.Plant Cell Physiol, 49(6): 925-933. |
| 9 | Hanzal-Bayer M, Renault L, Roversi P, Wittinghofer A, Hillig R C.2002. The complex of Arl2-GTP and PDEσ: From structure to function. EMBO J, 21(9): 2095-2106. |
| 10 | Hirose T, Terao T.2004. A comprehensive expression analysis of the starch synthase gene family in rice (Oryza sativa L.).Planta, 220(1): 9-16. |
| 11 | Jin T Y, Li H, Guo T, Liu X L, Su N, Wu F Q.2010. Analysis of physiological and biochemical characteristics of six mutants with stable high percentage of chalkiness in rice grains.Acta Agron Sin, 36(1): 121-132. (in Chinese with English abstract) |
| 12 | Kang H G, Park S, Matsuoka M, An G.2005. White-core endosperm floury endosperm 4 in rice is generated by knockout mutations in the C4-type pyruvate orthophosphate dikinase gene (OsPPDKB).Plant J, 42(6): 901-911. |
| 13 | Kang H Q, Cheng H Y.2007. Study on chalkiness characters and endosperm structures of the main parents’ kernel of hybrid rice.Chin Agric Sci Bull, 23(4): 180-185. (in Chinese with English abstract) |
| 14 | Kawagoe Y, Kubo A, Satoh H, Takaiwa F, Nakamjura Y.2005. Roles of isoamylase and ADP-glucose pyrophosphorylase in starch granule synthesis in rice endosperm.Plant J, 42(2): 164-174. |
| 15 | Kawasaki T, Mizuno K, Baba T, Shimada H.1993. Molecular analysis of the gene encoding a rice starch branching enzyme.Mol Gen Genet, 237(1/2): 10-16. |
| 16 | Li Z, Trick H N.2005. Rapid method for high-quality RNA isolation from seed endosperm containing high levels of starch.Bio-Techniques, 38(6): 872-876. |
| 17 | Liu F, Ren Y L, Wang Y H, Peng C, Zhou K N, Lv J, Guo X P, Zhang X, Zhong M S, Zhao S L, Jiang L, Wang H Y, Bao Y Q, Wan J M.2013. OsVPS9A functions cooperatively with OsRAB5A to regulate post-Golgi dense vesicle-mediated storage protein trafficking to the protein storage vacuole in rice endosperm cells.Mol Plant, 6(6): 1918-1932. |
| 18 | Myers A M, Morell M K, James M G, Ball S G.2000. Recent progress toward understanding biosynthesis of the amylopectin crystal.Plant Physiol, 122(4): 989-998. |
| 19 | Nishi A, Nakamura Y, Tanaka N, Satoh H.2001. Biochemical and genetic analysis of the effects of amylose-extender mutation in rice endosperm.Plant Physiol, 127(2): 459-472. |
| 20 | Nishio T, Iida S.1993. Mutants having a low content of 16-kDa allergenic protein in rice (Oryza sativa L.).Theor Appl Genet, 86(2/3): 317-321. |
| 21 | Peng C, Wang Y H, Liu F, Ren Y L, Zhou K N, Lv J, Zheng M, Zhao S L, Zhang L, Wang C M, Jiang L, Zhang X, Guo X P, Bao Y Q, Wan J M.2014. FLOURY ENDOSPERM6 encodes a CBM48 domain-containing protein involved in compound granule formation and starch synthesis in rice endosperm.Plant J, 77(6): 917-930. |
| 22 | Ren Y L, Wang Y H, Liu F, Zhou K N, Ding Y, Zhou F, Wang Y, Liu K, Gan L, Ma W W, Han X H, Zhang X, Guo X P, Wu F Q, Cheng Z J, Wang J L, Lei C L, Lin Q B, Jiang L, Wu C Y, Bao Y Q, Wang H Y, Wan J M.2014. GLUTELIN PRECURSOR ACCUMULATION3 encodes a regulator of post-Golgi vesicular traffic essential for vacuolar protein sorting in rice endosperm.Plant Cell, 26(1): 410-425. |
| 23 | Qu Y, Jin Z X, Liu H Y, Xu Z H, Zhu L N, Zheng G L, Zhu F X, Zhang Z C.2014. Analysis of expression characters of soluble starch synthase and isoform genes involved in japonica hybrid progeny.Chin J Rice Sci, 28(1): 23-31. (in Chinese with English abstract) |
| 24 | Ryoo N, Yu C, Park C S, Baik M Y, Park I M, Cho M H, Bhoo S H, An G, Hahn T R, Jeon J S.2007. Knockout of a starch synthase gene OsSSIIIa/Flo5 causes white-core floury endosperm in rice (Oryza sativa L.).Plant Cell Rep, 26(7): 1083-1095. |
| 25 | Satoh H, Omura T.1981. New endosperm mutations induced by chemical mutagens in rice Oryza sativa L.Jpn J Breeding, 31(3): 316-326. |
| 26 | Schmittgen T D, Livak K J.2008. Analyzing real-time PCR data by the comparative CT method.Nat Protoc, 3: 1101-1108. |
| 27 | She K C, Kusano H, Koizumi K, Yamakawa H, Hakata M, Imamura T, Fukuda M, Naito N, Tsurumaki Y, Yaeshima M, Tsuqe T, Matsumoto K, Kudoh M, Itoh E, Kikuchi S, Kishimoto N, Yazaki J, Ando T, Yano M, Aoyama T, Sasaki T, Satoh H, Shimada H.2010. A novel factor FLOURY ENDOSPERM2 is involved in regulation of rice grain size and starch quality.Plant Cell, 22(10): 3280-3294. |
| 28 | Shimada H, Tada Y, Kawasaki T, Fujimura T.1993. Antisense regulation of the rice waxy gene expression using a PCR-amplified fragment of the rice genome reduces the amylose content in grain starch.Theor Appl Genet, 86(6): 665-672. |
| 29 | Steegmans M, Iliaens S, Hoebregs H.2004. Enzymatic, spectro- photometric determination of glucose, fructose, sucrose, and inulin/oligofructose in foods. J AOAC Int, 87(5): 1200-1207. |
| 30 | Sun C X, Duan B W, Xie L H, Chen N.2006. Determination of several quality characteristics of brown rice by near infrared transmission spectroscopy.Chin J Rice Sci, 20(4): 451-454. (in Chinese with English abstract) |
| 31 | Wang J C, Xu H, Zhu Y, Liu Q Q, Cai X L.2013.OsbZIP58, a basic leucine zipper transcription factor, regulates starch biosynthesis in rice endosperm.J Exp Bot, 64(11): 3453-3466. |
| 32 | Wang Y H, Ren Y L, Liu X, Jiang L, Chen L M, Han X H, Jin M N, Liu S J, Liu F, Lv J, Zhou K N, Su N, Bao Y Q, Wan J M.2010. OsRab5a regulates endomembrane organization and storage protein trafficking in rice endosperm cells. Plant J, 64(5): 812-824. |
| 33 | Woo M O, Ham T H, Ji H S, Choi M S, Jiang W Z, Chu S H, Piao R H, Chin J H, Kim J A, Park B S, Seo H S, Jwa N S, McCouch S, Koh H J.2008. Inactivation of the UGPase1 gene causes genic male sterility and endosperm chalkiness in rice (Oryza sativa L.). Plant J, 54(2): 190-204. |
| 34 | Wu K S, Tanksley S D.1993. Abundance, polymorphism and genetic mapping of microsatellites in rice.Mol Gen Genet, 241(1/2): 225-235. |
| 35 | Yao D S, Song R T.2013. Research progress of maize opaque endosperm mutants.Chin J Nat, 35(2): 105-111. (in Chinese with English abstract) |
| 36 | (Managing Editor: Li Guan) |
| [1] | LI Qianlong, FENG Qi, WANG Heqin, KANG Yunhai, ZHANG Conghe, DU Ming, ZHANG Yunhu, WANG Hui, CHEN Jinjie, HAN Bin, FANG Yu, WANG Ahong. Genome-Wide Dissection of Quan 9311A Breeding Process and Application Advantages [J]. Rice Science, 2023, 30(6): 7-. |
| [2] | JI Dongling, XIAO Wenhui, SUN Zhiwei, LIU Lijun, GU Junfei, ZHANG Hao, Tom Matthew HARRISON, LIU Ke, WANG Zhiqin, WANG Weilu, YANG Jianchang. Translocation and Distribution of Carbon-Nitrogen in Relation to Rice Yield and Grain Quality as Affected by High Temperature at Early Panicle Initiation Stage [J]. Rice Science, 2023, 30(6): 12-. |
| [3] | Prathap V, Suresh KUMAR, Nand Lal MEENA, Chirag MAHESHWARI, Monika DALAL, Aruna TYAGI. Phosphorus Starvation Tolerance in Rice Through a Combined Physiological, Biochemical and Proteome Analysis [J]. Rice Science, 2023, 30(6): 8-. |
| [4] | Serena REGGI, Elisabetta ONELLI, Alessandra MOSCATELLI, Nadia STROPPA, Matteo Dell’ANNO, Kiril PERFANOV, Luciana ROSSI. Seed-Specific Expression of Apolipoprotein A-IMilano Dimer in Rice Engineered Lines [J]. Rice Science, 2023, 30(6): 6-. |
| [5] | Sundus ZAFAR, XU Jianlong. Recent Advances to Enhance Nutritional Quality of Rice [J]. Rice Science, 2023, 30(6): 4-. |
| [6] | Kankunlanach KHAMPUANG, Nanthana CHAIWONG, Atilla YAZICI, Baris DEMIRER, Ismail CAKMAK, Chanakan PROM-U-THAI. Effect of Sulfur Fertilization on Productivity and Grain Zinc Yield of Rice Grown under Low and Adequate Soil Zinc Applications [J]. Rice Science, 2023, 30(6): 9-. |
| [7] | FAN Fengfeng, CAI Meng, LUO Xiong, LIU Manman, YUAN Huanran, CHENG Mingxing, Ayaz AHMAD, LI Nengwu, LI Shaoqing. Novel QTLs from Wild Rice Oryza longistaminata Confer Rice Strong Tolerance to High Temperature at Seedling Stage [J]. Rice Science, 2023, 30(6): 14-. |
| [8] | XIA Xiaodong, ZHANG Xiaobo, WANG Zhonghao, CHENG Benyi, Sun Huifeng, XU Xia, GONG Junyi, YANG Shihua, WU Jianli, SHI Yongfeng, XU Rugen. Mapping and Functional Analysis of LE Gene in a Lethal Etiolated Rice Mutant at Seedling Stage [J]. Rice Science, 2023, 30(6): 13-. |
| [9] | LIN Shaodan, YAO Yue, LI Jiayi, LI Xiaobin, MA Jie, WENG Haiyong, CHENG Zuxin, YE Dapeng. Application of UAV-Based Imaging and Deep Learning in Assessment of Rice Blast Resistance [J]. Rice Science, 2023, 30(6): 10-. |
| [10] | Md. Forshed DEWAN, Md. AHIDUZZAMAN, Md. Nahidul ISLAM, Habibul Bari SHOZIB. Potential Benefits of Bioactive Compounds of Traditional Rice Grown in South and South-East Asia: A Review [J]. Rice Science, 2023, 30(6): 5-. |
| [11] | Raja CHAKRABORTY, Pratap KALITA, Saikat SEN. Phenolic Profile, Antioxidant, Antihyperlipidemic and Cardiac Risk Preventive Effect of Chakhao Poireiton (A Pigmented Black Rice) in High-Fat High-Sugar induced Rats [J]. Rice Science, 2023, 30(6): 11-. |
| [12] | Nazaratul Ashifa Abdullah Salim, Norlida Mat Daud, Julieta Griboff, Abdul Rahim Harun. Elemental Assessments in Paddy Soil for Geographical Traceability of Rice from Peninsular Malaysia [J]. Rice Science, 2023, 30(5): 486-498. |
| [13] | Monica Ruffini Castiglione, Stefania Bottega, Carlo Sorce, Carmelina SpanÒ. Effects of Zinc Oxide Particles with Different Sizes on Root Development in Oryza sativa [J]. Rice Science, 2023, 30(5): 449-458. |
| [14] | Ammara Latif, Sun Ying, Pu Cuixia, Noman Ali. Rice Curled Its Leaves Either Adaxially or Abaxially to Combat Drought Stress [J]. Rice Science, 2023, 30(5): 405-416. |
| [15] | Liu Qiao, Qiu Linlin, Hua Yangguang, Li Jing, Pang Bo, Zhai Yufeng, Wang Dekai. LHD3 Encoding a J-Domain Protein Controls Heading Date in Rice [J]. Rice Science, 2023, 30(5): 437-448. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||